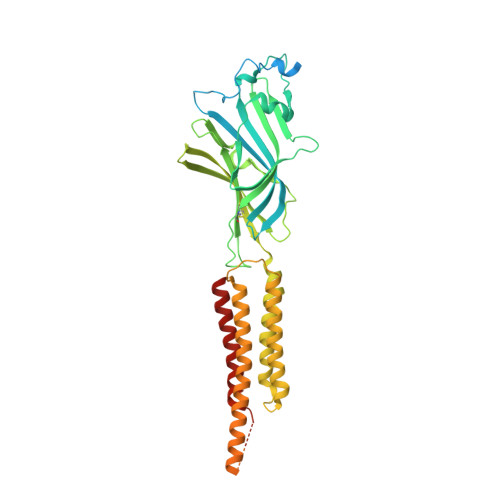
Structure and dynamics of differential ligand binding in the human rho-type GABA A receptor.
Cowgill, J., Fan, C., Haloi, N., Tobiasson, V., Zhuang, Y., Howard, R.J., Lindahl, E.(2023) Neuron 111: 3450
- PubMed: 37659407 Search on PubMed
- DOI: https://doi.org/10.1016/j.neuron.2023.08.006
- Primary Citation Related Structures:
8OP9, 8OQ6, 8OQ7, 8OQ8, 8OQA - PubMed Abstract:
The neurotransmitter γ-aminobutyric acid (GABA) drives critical inhibitory processes in and beyond the nervous system, partly via ionotropic type-A receptors (GABA A Rs). Pharmacological properties of ρ-type GABA A Rs are particularly distinctive, yet the structural basis for their specialization remains unclear. Here, we present cryo-EM structures of a lipid-embedded human ρ1 GABA A R, including a partial intracellular domain, under apo, inhibited, and desensitized conditions. An apparent resting state, determined first in the absence of modulators, was recapitulated with the specific inhibitor (1,2,5,6-tetrahydropyridin-4-yl)methylphosphinic acid and blocker picrotoxin and provided a rationale for bicuculline insensitivity. Comparative structures, mutant recordings, and molecular simulations with and without GABA further explained the sensitized but slower activation of ρ1 relative to canonical subtypes. Combining GABA with picrotoxin also captured an apparent uncoupled intermediate state. This work reveals structural mechanisms of gating and modulation with applications to ρ-specific pharmaceutical design and to our biophysical understanding of ligand-gated ion channels.
- Department of Biochemistry and Biophysics, SciLifeLab, Stockholm University, 17121 Solna, Sweden.
Organizational Affiliation: