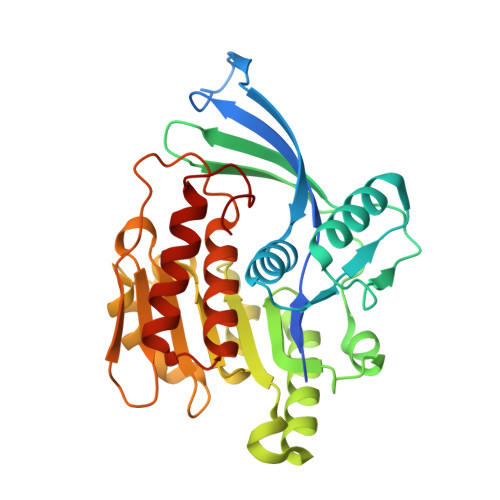
Crystal structures of human and mouse ketohexokinase provide a structural basis for species- and isoform-selective inhibitor design.
Ebenhoch, R., Bauer, M., Romig, H., Gottschling, D., Kley, J.T., Heine, N., Weber, A., Uphues, I., Nar, H., Pautsch, A.(2023) Acta Crystallogr D Struct Biol 79: 871-880
- PubMed: 37712434 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798323006137
- Primary Citation Related Structures:
8OMD, 8OME, 8OMF, 8OMG - PubMed Abstract:
A molecular understanding of the proteins involved in fructose metabolism is essential for controlling the current spread of fructose-related obesity, diabetes and related adverse metabolic states in Western populations. Fructose catabolism starts with the phosphorylation of D-fructose to fructose 1-phosphate by ketohexokinase (KHK). KHK exists in two alternatively spliced isoforms: the hepatic and intestinal isoform KHK-C and the peripheral isoform KHK-A. Here, the structure of apo murine KHK (mKHK), which differs from structures of human KHK in overall conformation, is reported. An isoform-selective ligand, which offers a 50-fold higher potency on mKHK and human KHK-A compared with KHK-C, is further characterized. In mKHK, large-scale conformational changes are observed upon ligand binding. The structures suggest a combined strategy for the design of species- and isoform-selective KHK inhibitors.
- Medicinal Chemistry, Boehringer Ingelheim Pharma GmbH & Co. KG, Birkendorferstrasse 67, 88400 Biberach an der Riss, Germany.
Organizational Affiliation: