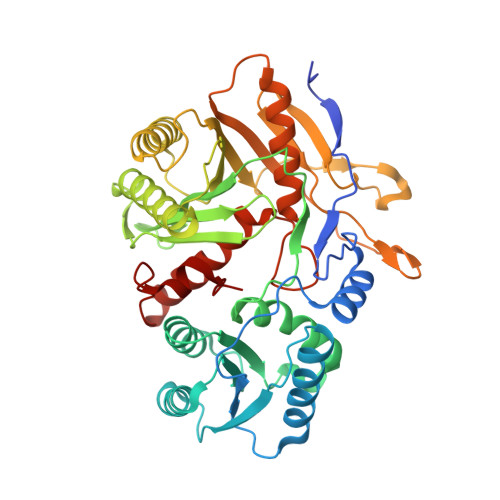
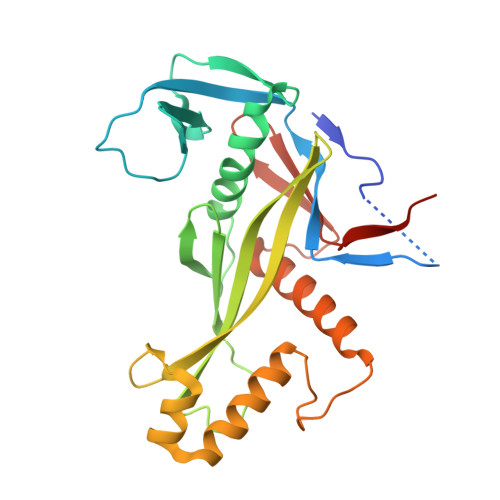
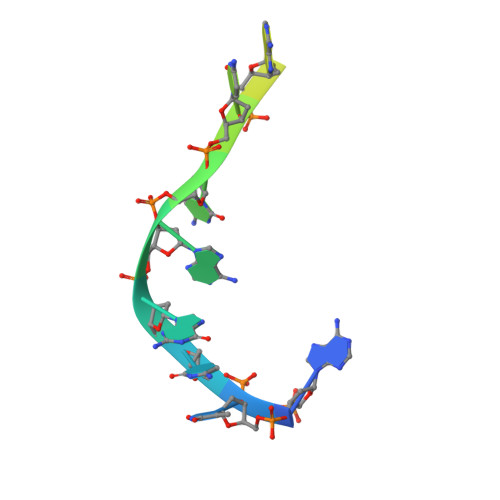
The missing part: the Archaeoglobus fulgidus Argonaute forms a functional heterodimer with an N-L1-L2 domain protein.
Manakova, E., Golovinas, E., Poceviciute, R., Sasnauskas, G., Silanskas, A., Rutkauskas, D., Jankunec, M., Zagorskaite, E., Jurgelaitis, E., Grybauskas, A., Venclovas, C., Zaremba, M.(2024) Nucleic Acids Res 52: 2530-2545
- PubMed: 38197228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkad1241
- Primary Citation Related Structures:
8OK9, 8OLD, 8OLJ, 8PVV, 8QG0 - PubMed Abstract:
Argonaute (Ago) proteins are present in all three domains of life (bacteria, archaea and eukaryotes). They use small (15-30 nucleotides) oligonucleotide guides to bind complementary nucleic acid targets and are responsible for gene expression regulation, mobile genome element silencing, and defence against viruses or plasmids. According to their domain organization, Agos are divided into long and short Agos. Long Agos found in prokaryotes (long-A and long-B pAgos) and eukaryotes (eAgos) comprise four major functional domains (N, PAZ, MID and PIWI) and two structural linker domains L1 and L2. The majority (∼60%) of pAgos are short pAgos, containing only the MID and inactive PIWI domains. Here we focus on the prokaryotic Argonaute AfAgo from Archaeoglobus fulgidus DSM4304. Although phylogenetically classified as a long-B pAgo, AfAgo contains only MID and catalytically inactive PIWI domains, akin to short pAgos. We show that AfAgo forms a heterodimeric complex with a protein encoded upstream in the same operon, which is a structural equivalent of the N-L1-L2 domains of long pAgos. This complex, structurally equivalent to a long PAZ-less pAgo, outperforms standalone AfAgo in guide RNA-mediated target DNA binding. Our findings provide a missing piece to one of the first and the most studied pAgos.
- Institute of Biotechnology, Life Sciences Center, Vilnius University, Sauletekio av. 7, LT-10257, Vilnius, Lithuania.
Organizational Affiliation: