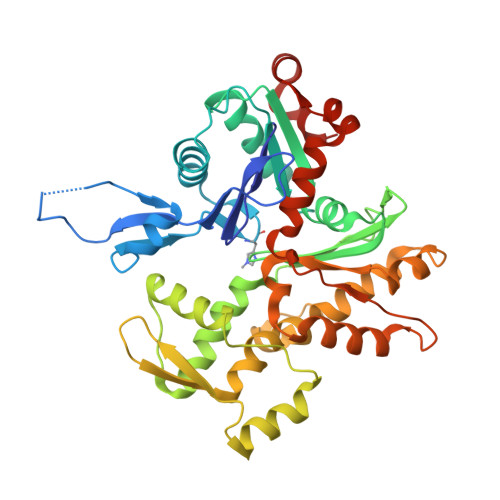
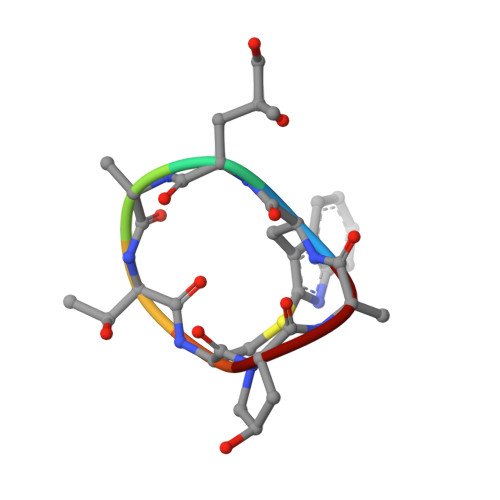
Molecular mechanisms of inorganic-phosphate release from the core and barbed end of actin filaments.
Oosterheert, W., Blanc, F.E.C., Roy, A., Belyy, A., Sanders, M.B., Hofnagel, O., Hummer, G., Bieling, P., Raunser, S.(2023) Nat Struct Mol Biol 30: 1774-1785
- PubMed: 37749275 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01101-9
- Primary Citation Related Structures:
8OI6, 8OI8, 8OID - PubMed Abstract:
The release of inorganic phosphate (P i ) from actin filaments constitutes a key step in their regulated turnover, which is fundamental to many cellular functions. The mechanisms underlying P i release from the core and barbed end of actin filaments remain unclear. Here, using human and bovine actin isoforms, we combine cryo-EM with molecular-dynamics simulations and in vitro reconstitution to demonstrate how actin releases P i through a 'molecular backdoor'. While constantly open at the barbed end, the backdoor is predominantly closed in filament-core subunits and opens only transiently through concerted amino acid rearrangements. This explains why P i escapes rapidly from the filament end but slowly from internal subunits. In a nemaline-myopathy-associated actin variant, the backdoor is predominantly open in filament-core subunits, resulting in accelerated P i release and filaments with drastically shortened ADP-P i caps. Our results provide the molecular basis for P i release from actin and exemplify how a disease-linked mutation distorts the nucleotide-state distribution and atomic structure of the filament.
- Department of Structural Biochemistry, Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: