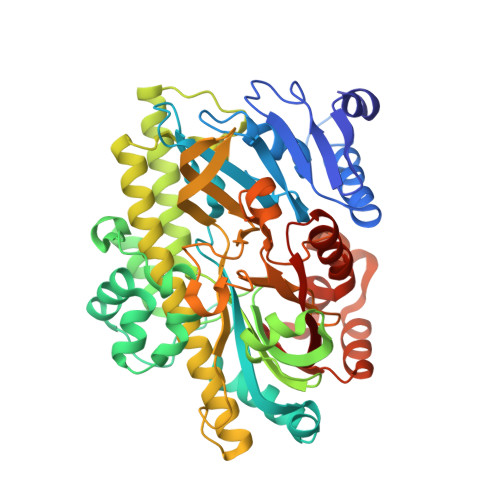
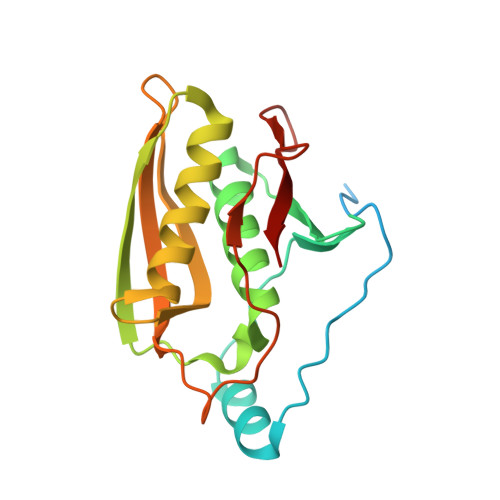
Structural and mechanistic insights into activation of the human RNA ligase RTCB by Archease.
Gerber, J.L., Morales Guzman, S.I., Worf, L., Hubbe, P., Kopp, J., Peschek, J.(2024) Nat Commun 15: 2378-2378
- PubMed: 38493148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46568-2
- Primary Citation Related Structures:
8BTT, 8BTX, 8ODO, 8ODP - PubMed Abstract:
RNA ligases of the RTCB-type play an essential role in tRNA splicing, the unfolded protein response and RNA repair. RTCB is the catalytic subunit of the pentameric human tRNA ligase complex. RNA ligation by the tRNA ligase complex requires GTP-dependent activation of RTCB. This active site guanylylation reaction relies on the activation factor Archease. The mechanistic interplay between both proteins has remained unknown. Here, we report a biochemical and structural analysis of the human RTCB-Archease complex in the pre- and post-activation state. Archease reaches into the active site of RTCB and promotes the formation of a covalent RTCB-GMP intermediate through coordination of GTP and metal ions. During the activation reaction, Archease prevents futile RNA substrate binding to RTCB. Moreover, monomer structures of Archease and RTCB reveal additional states within the RNA ligation mechanism. Taken together, we present structural snapshots along the reaction cycle of the human tRNA ligase.
- Heidelberg University, Biochemistry Center (BZH), Im Neuenheimer Feld 328, Heidelberg, Germany.
Organizational Affiliation: