The CENP-O complex links inner kinetochore to centromere cohesion
Liu, M.J., He, X.J.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cohesin subunit SA-2 | 980 | Homo sapiens | Mutation(s): 0 Gene Names: SA2 | 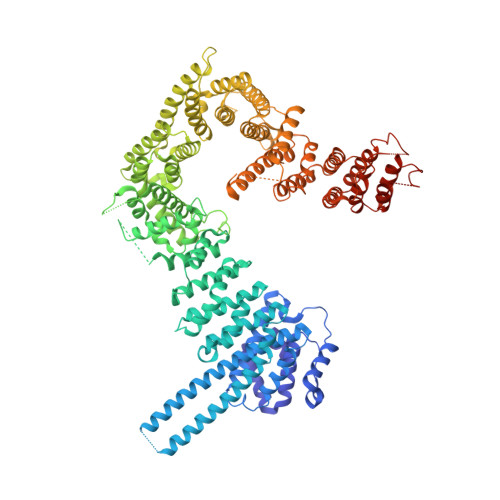 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q8N3U4 GTEx: ENSG00000101972 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8N3U4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 64-kDa C-terminal product | 141 | Homo sapiens | Mutation(s): 0 Gene Names: SCC1 | 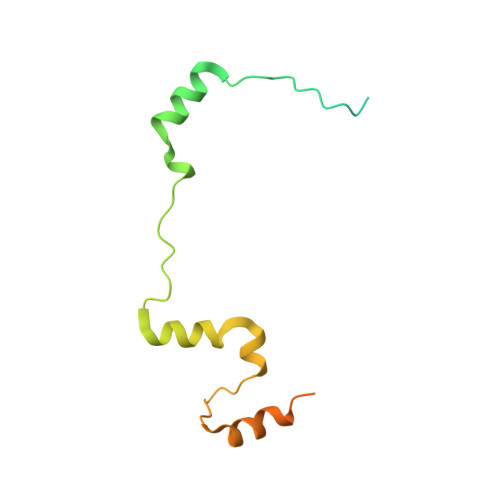 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O60216 GTEx: ENSG00000164754 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O60216 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CENP-U | 9 | Homo sapiens | Mutation(s): 0 | 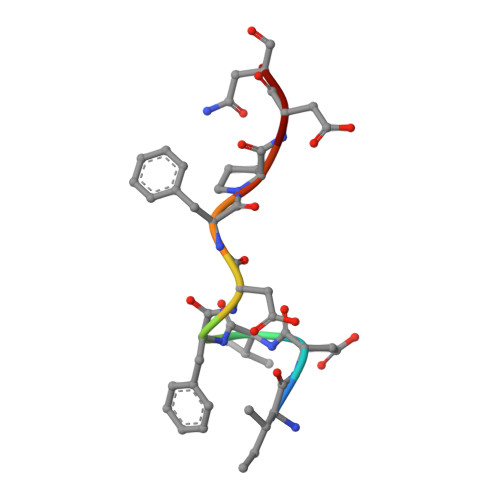 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q71F23 GTEx: ENSG00000151725 | |||||
Entity Groups | |||||
| UniProt Group | Q71F23 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 79.321 | α = 90 |
| b = 132.615 | β = 90 |
| c = 148.001 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |