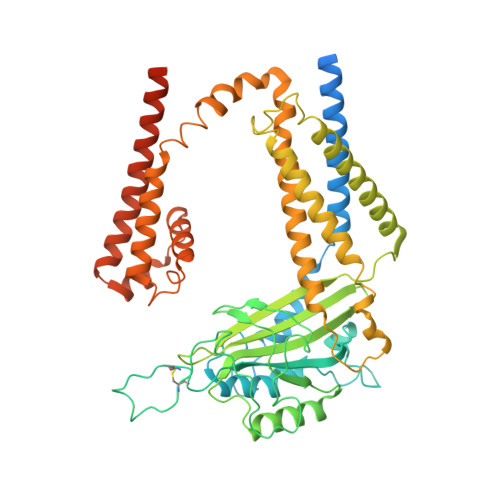
Molecular and structural basis of the dual regulation of the polycystin-2 ion channel by small-molecule ligands.
Wang, Z., Chen, M., Su, Q., Morais, T.D.C., Wang, Y., Nazginov, E., Pillai, A.R., Qian, F., Shi, Y., Yu, Y.(2024) Proc Natl Acad Sci U S A 121: e2316230121-e2316230121
- PubMed: 38483987 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2316230121
- Primary Citation Related Structures:
8HK7, 8K3S - PubMed Abstract:
Mutations in the PKD2 gene, which encodes the polycystin-2 (PC2, also called TRPP2) protein, lead to autosomal dominant polycystic kidney disease (ADPKD). As a member of the transient receptor potential (TRP) channel superfamily, PC2 functions as a non-selective cation channel. The activation and regulation of the PC2 channel are largely unknown, and direct binding of small-molecule ligands to this channel has not been reported. In this work, we found that most known small-molecule agonists of the mucolipin TRP (TRPML) channels inhibit the activity of the PC2_F604P, a gain-of-function mutant of the PC2 channel. However, two of them, ML-SA1 and SF-51, have dual regulatory effects, with low concentration further activating PC2_F604P, and high concentration leading to inactivation of the channel. With two cryo-electron microscopy (cryo-EM) structures, a molecular docking model, and mutagenesis results, we identified two distinct binding sites of ML-SA1 in PC2_F604P that are responsible for activation and inactivation, respectively. These results provide structural and functional insights into how ligands regulate PC2 channel function through unusual mechanisms and may help design compounds that are more efficient and specific in regulating the PC2 channel and potentially also for ADPKD treatment.
- Department of Biological Sciences, St. John's University, Queens, NY 11375.
Organizational Affiliation: