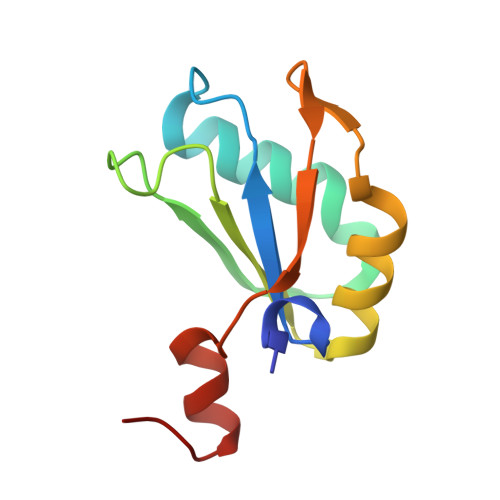
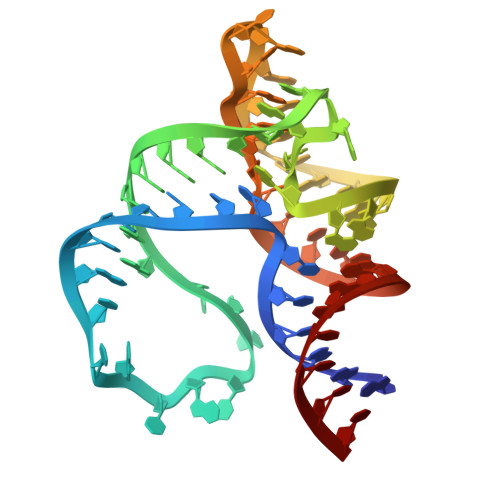
Structural mechanisms for binding and activation of a contact-quenched fluorophore by RhoBAST.
Zhang, Y., Xu, Z., Xiao, Y., Jiang, H., Zuo, X., Li, X., Fang, X.(2024) Nat Commun 15: 4206-4206
- PubMed: 38760339 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-48478-9
- Primary Citation Related Structures:
8JY0 - PubMed Abstract:
The fluorescent light-up aptamer RhoBAST, which binds and activates the fluorophore-quencher conjugate tetramethylrhodamine-dinitroaniline with high affinity, super high brightness, remarkable photostability, and fast exchange kinetics, exhibits excellent performance in super-resolution RNA imaging. Here we determine the co-crystal structure of RhoBAST in complex with tetramethylrhodamine-dinitroaniline to elucidate the molecular basis for ligand binding and fluorescence activation. The structure exhibits an asymmetric "A"-like architecture for RhoBAST with a semi-open binding pocket harboring the xanthene of tetramethylrhodamine at the tip, while the dinitroaniline quencher stacks over the phenyl of tetramethylrhodamine instead of being fully released. Molecular dynamics simulations show highly heterogeneous conformational ensembles with the contact-but-unstacked fluorophore-quencher conformation for both free and bound tetramethylrhodamine-dinitroaniline being predominant. The simulations also show that, upon RNA binding, the fraction of xanthene-dinitroaniline stacked conformation significantly decreases in free tetramethylrhodamine-dinitroaniline. This highlights the importance of releasing dinitroaniline from xanthene tetramethylrhodamine to unquench the RhoBAST-tetramethylrhodamine-dinitroaniline complex. Using SAXS and ITC, we characterized the magnesium dependency of the folding and binding mode of RhoBAST in solution and indicated its strong structural robustness. The structures and binding modes of relevant fluorescent light-up aptamers are compared, providing mechanistic insights for rational design and optimization of this important fluorescent light-up aptamer-ligand system.
- Key Laboratory of RNA Science and Engineering, Institute of Biophysics Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: