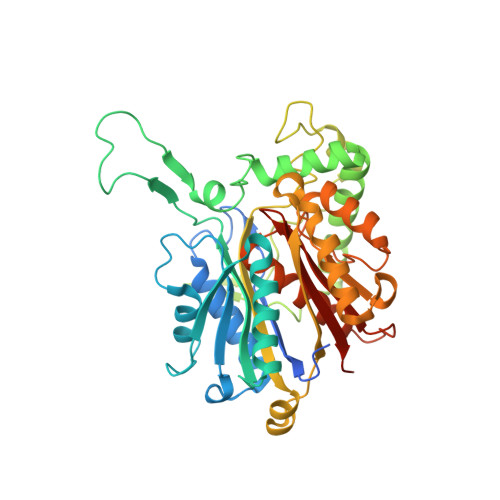
Molecular Chain Elongation Mechanism for n-Caproate Biosynthesis by Megasphaera Hexanoica.
Jeon, B.S., Kim, E.J., Seo, H., Kim, H., Shin, S., Schlaiss, C., Angenent, L.T., Kim, K.J., Sang, B.I.(2025) Adv Sci (Weinh) : e06069-e06069
- PubMed: 40932660 Search on PubMed
- DOI: https://doi.org/10.1002/advs.202506069
- Primary Citation Related Structures:
8JG2, 8JG3 - PubMed Abstract:
The microbial production of medium-chain carboxylates has attracted considerable interest owing to their potential applications in biofuels and specialty chemicals; however, the underlying biosynthetic mechanisms remain incompletely understood. The present study evaluates the medium-chain carboxylate-producing microbe Megaspahera hexanoica using genomic analysis, transcriptome analysis, and metabolic engineering. Additionally, the n-caproate synthesis pathway of M. hexanoica is characterized with fructose as an electron donor, and the substrate specificity of the respective proteins is evaluated by constructing an n-caproate biosynthetic pathway in Escherichia coli. Among all r-BOX or RBO genes, thl_1583, which encodes β-ketothiolase (MhTHL), is identified as the most critical enzyme for the carbon chain elongation mechanism in M. hexanoica. Therefore, MhTHL is compared with other well-studied β-ketothiolases (CkTHL from Clostridium kluyveri, ReBktB from Ralstonia eutropha (Cupriavidus necator), EcAtoB from E. coli, and CaTHL from C. acetobutylicum). MhTHL is found to exhibit the highest n-caproate production, as evidenced by the protein crystal structure of MhTHL. Structural comparisons with other thiolases show that MhTHL has a larger substrate-binding pocket than ReBktB. Thiolase mutants generated by site-directed mutagenesis reveal that two residues (Leu87 and Val351) are essential for determining the size of the substrate-binding pocket.
- Korea Institute of Ceramic Engineering and Technology, Osong, 28160, Republic of Korea.
Organizational Affiliation: