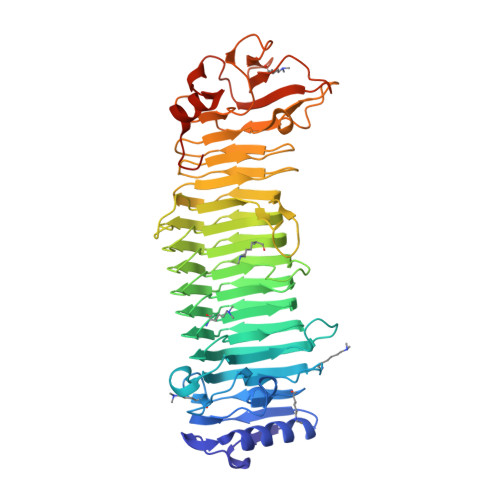
Structural basis for the minimal bifunctional alginate epimerase AlgE3 from Azotobacter chroococcum.
Fujiwara, T., Mano, E., Nango, E.(2024) FEBS Lett 598: 1422-1437
- PubMed: 38649293 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.14886
- Primary Citation Related Structures:
8JA4, 8JA6, 8JAZ, 8XFQ, 8XFR - PubMed Abstract:
Among the epimerases specific to alginate, some of them in Azotobacter genera convert β-d-mannuronic acid to α-l-guluronic acid but also have lyase activity to degrade alginate. The remarkable characteristics of these epimerases make it a promising enzyme for tailoring alginates to meet specific demands. Here, we determined the structure of the bifunctional mannuronan C-5 epimerase AlgE3 from Azotobacter chroococcum (AcAlgE3) in complex with several mannuronic acid oligomers as well as in apo form, which allowed us to elucidate the binding manner of each mannuronic acid oligomer, and the structural plasticity, which is dependent on calcium ions. Moreover, a comprehensive analysis of the lyase activity profiles of AcAlgE3 combined with structural characteristics explained the preference for different chain length oligomers.
- Institute of Multidisciplinary Research for Advanced Materials, Tohoku University, Sendai, Japan.
Organizational Affiliation: