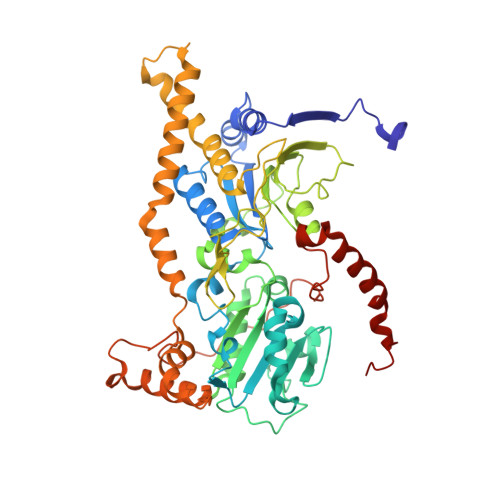
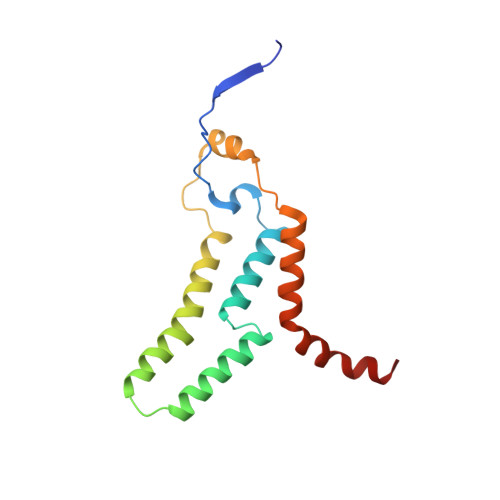
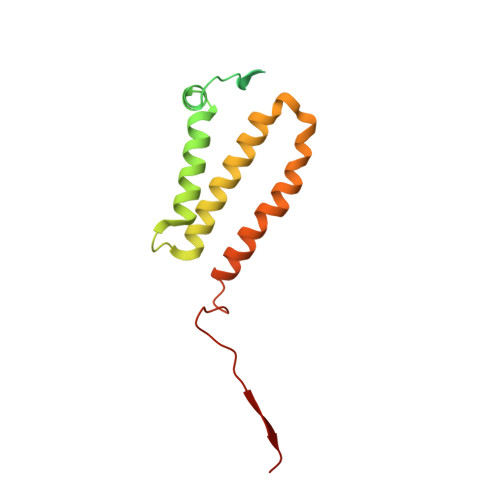
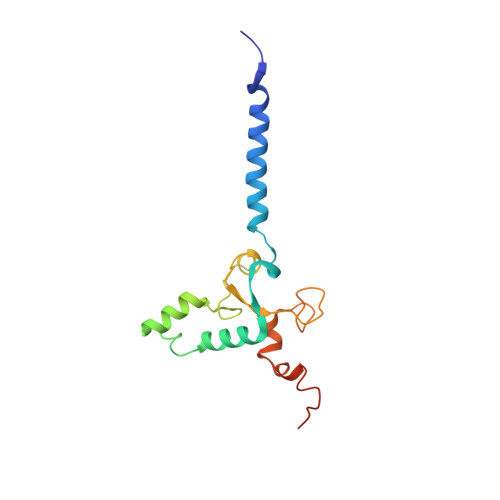
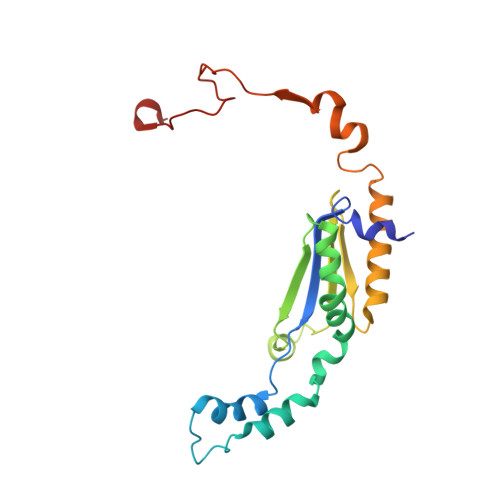
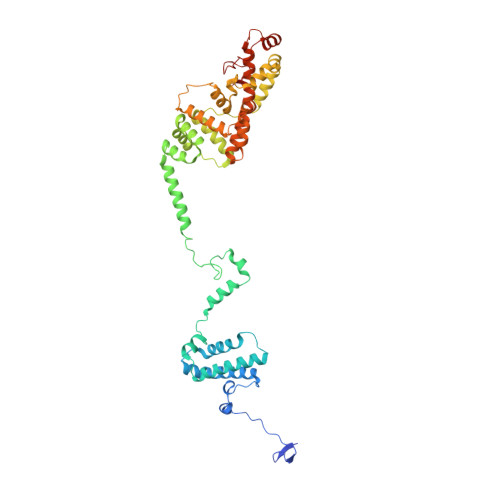
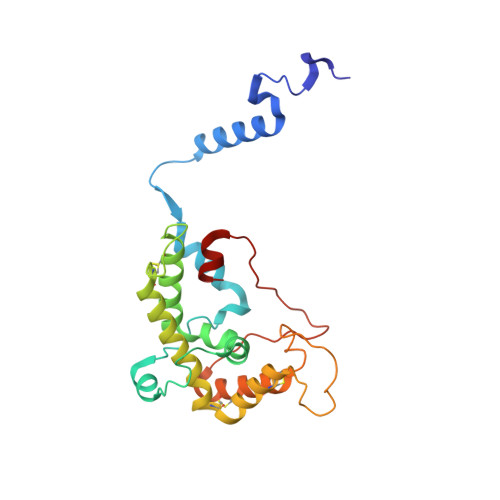
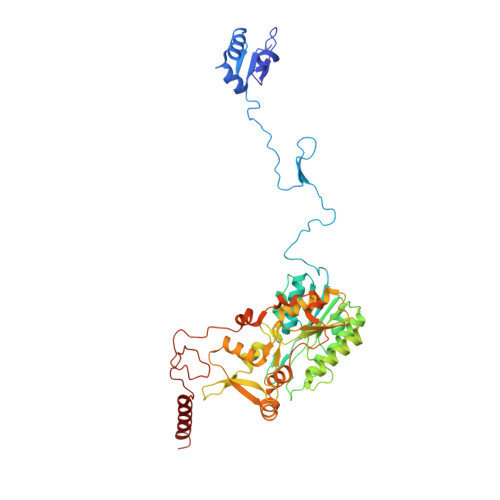
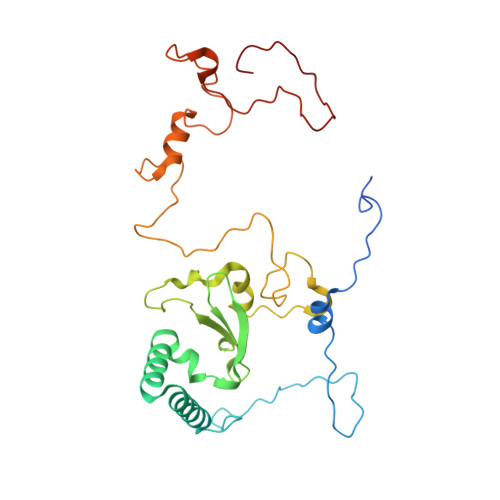
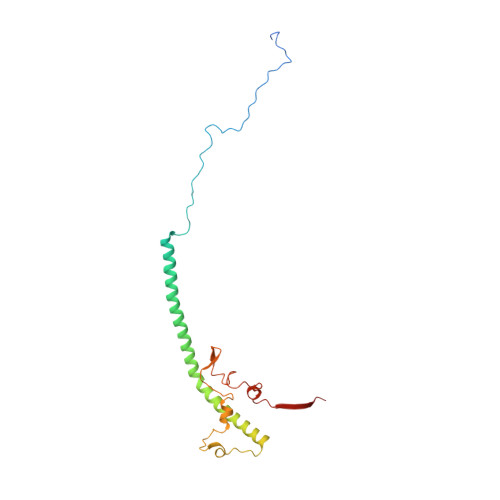
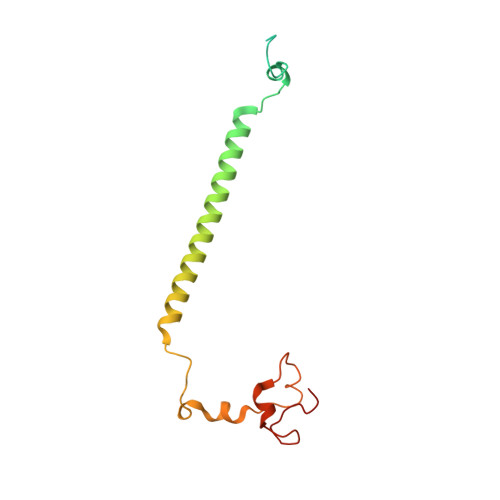
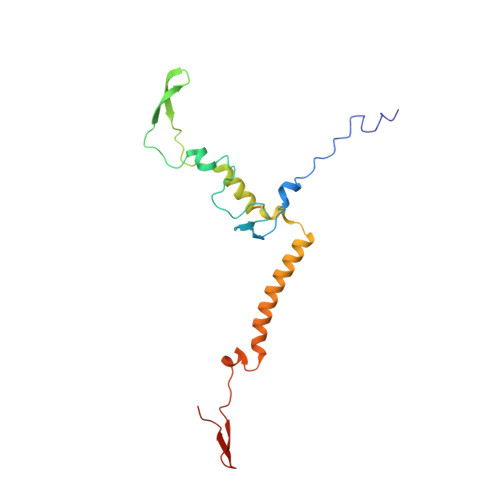
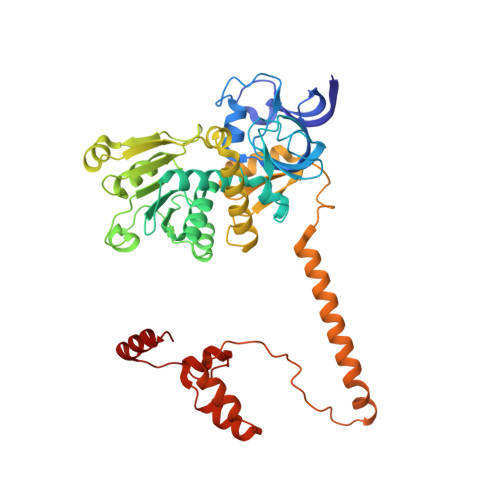
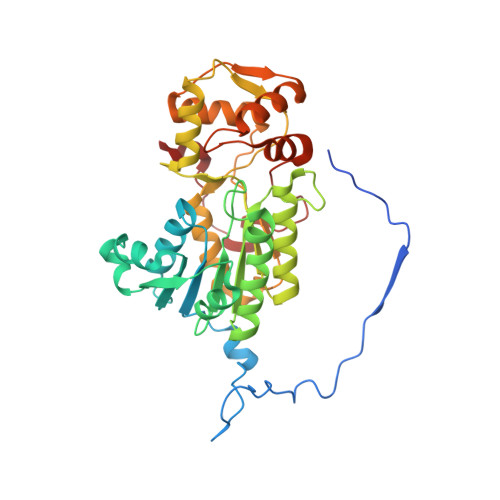
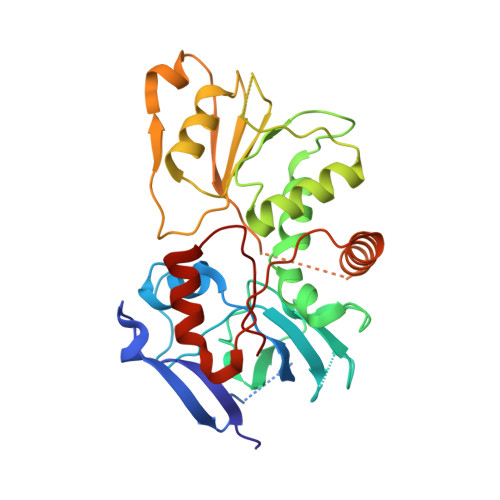
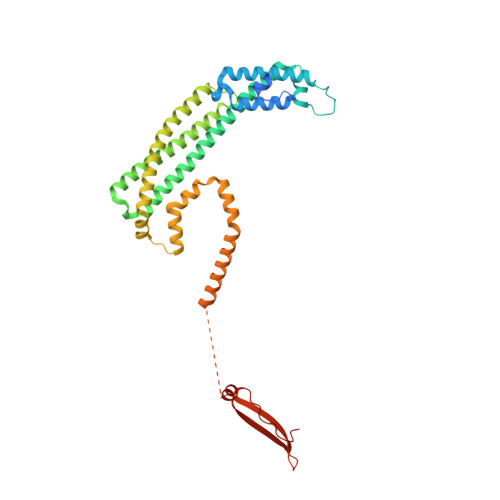
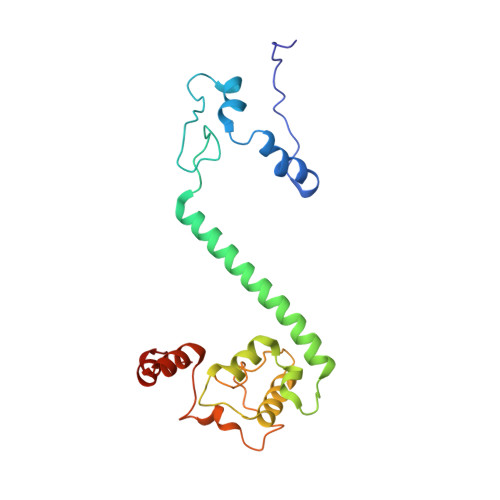
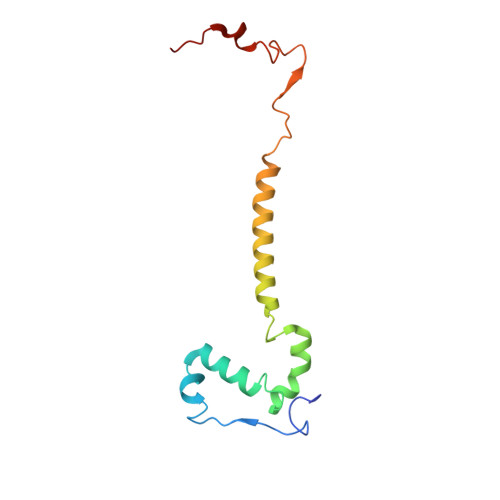
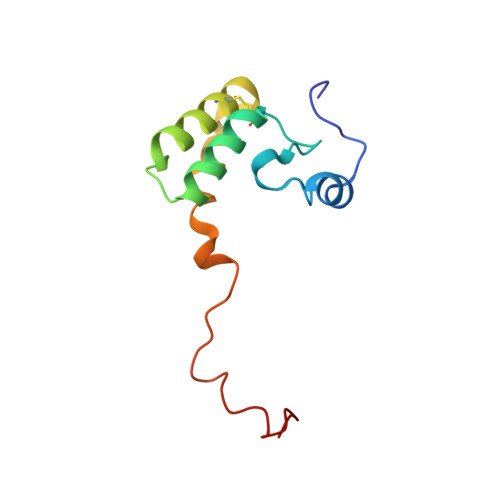
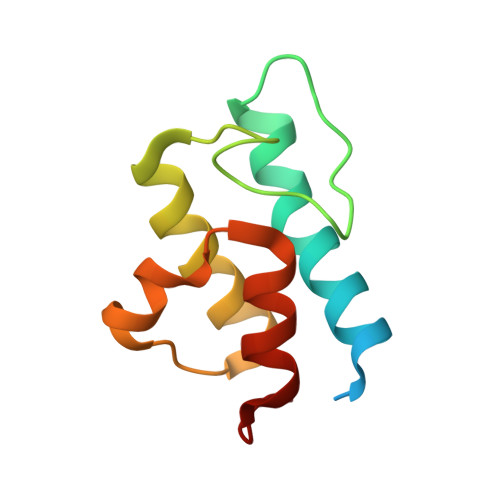
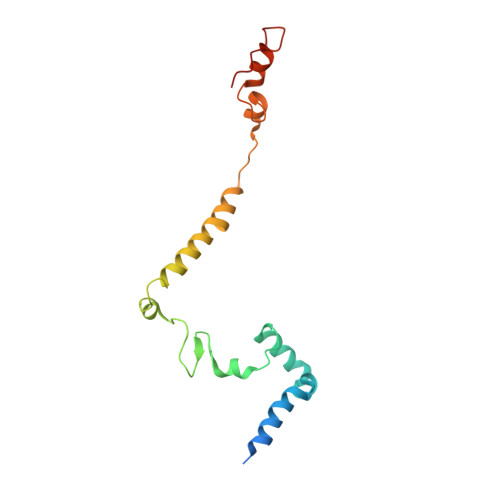
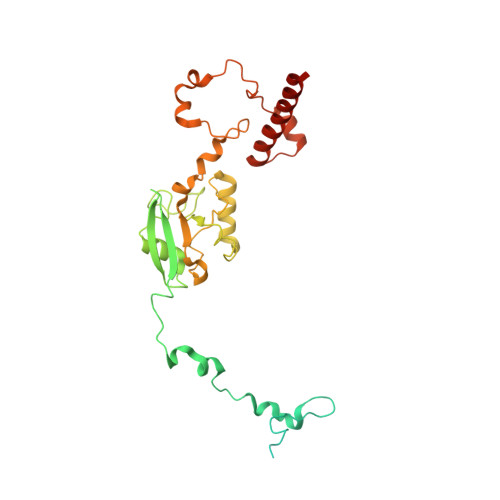
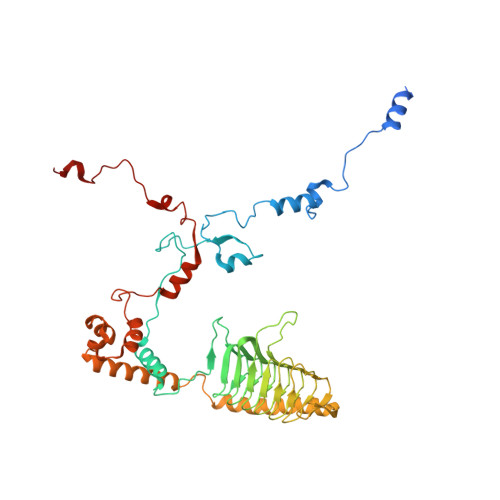
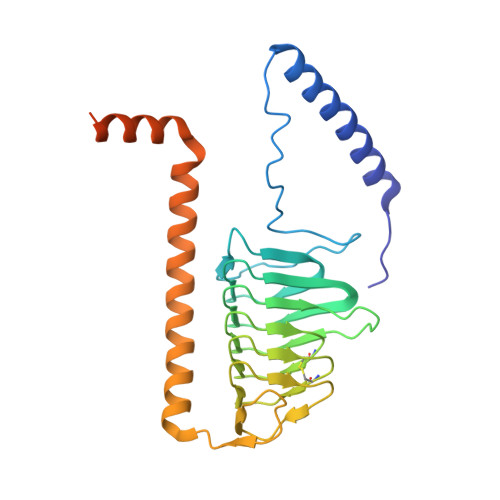
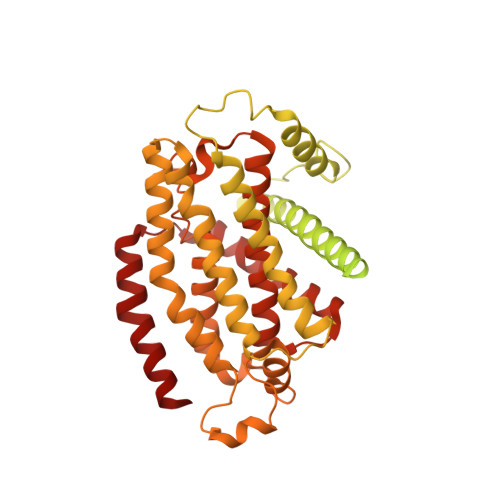
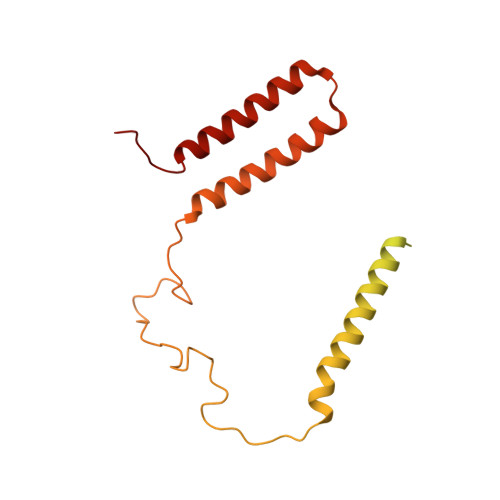
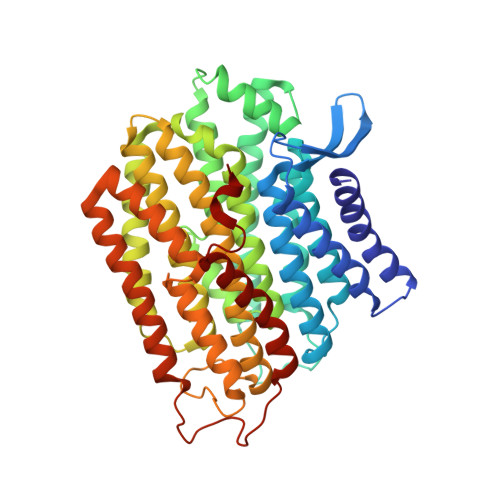
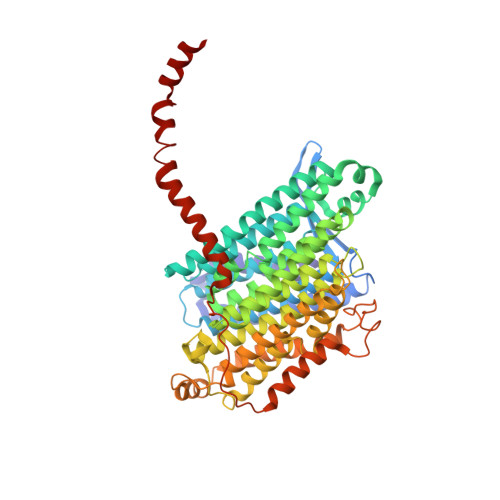
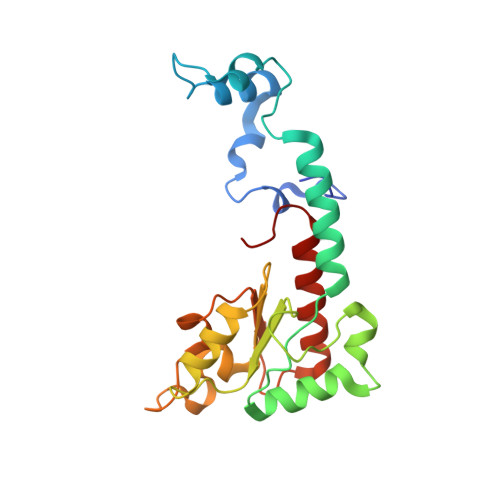
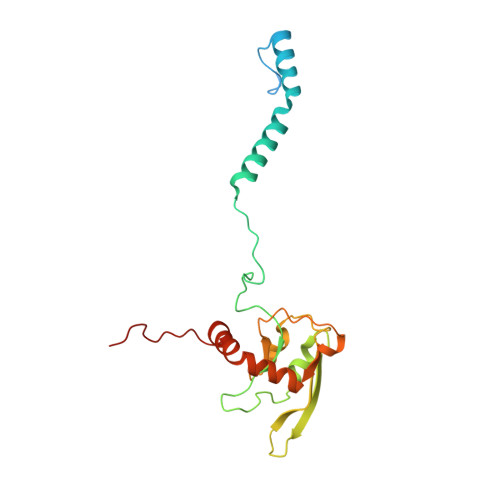
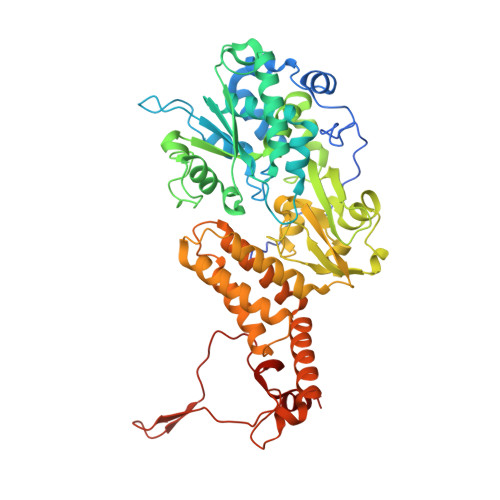
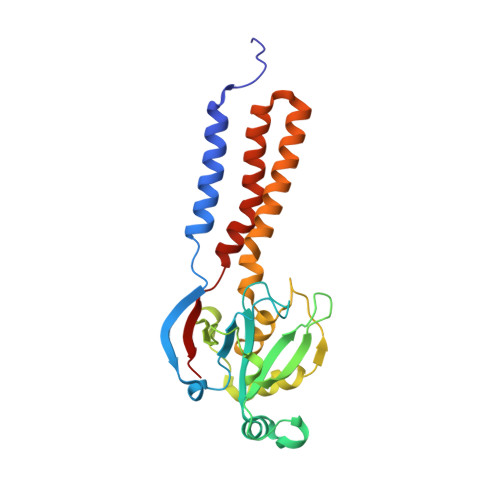
Euglena's atypical respiratory chain adapts to the discoidal cristae and flexible metabolism.
He, Z., Wu, M., Tian, H., Wang, L., Hu, Y., Han, F., Zhou, J., Wang, Y., Zhou, L.(2024) Nat Commun 15: 1628-1628
- PubMed: 38388527 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46018-z
- Primary Citation Related Structures:
8IUF, 8IUJ, 8J9H, 8J9I, 8J9J - PubMed Abstract:
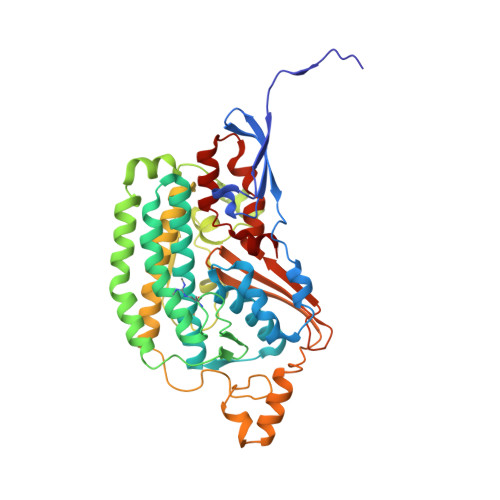
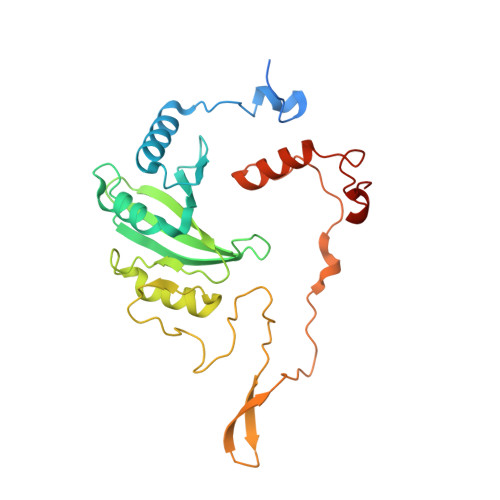
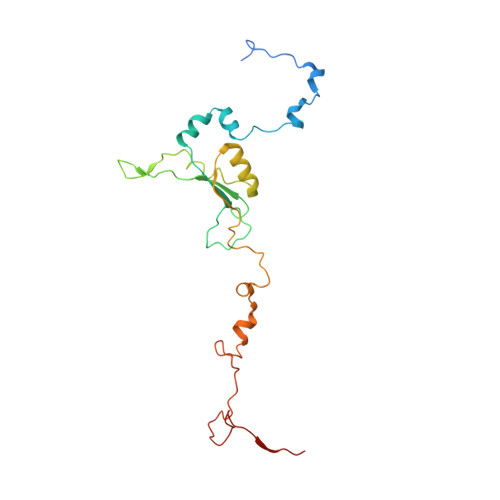
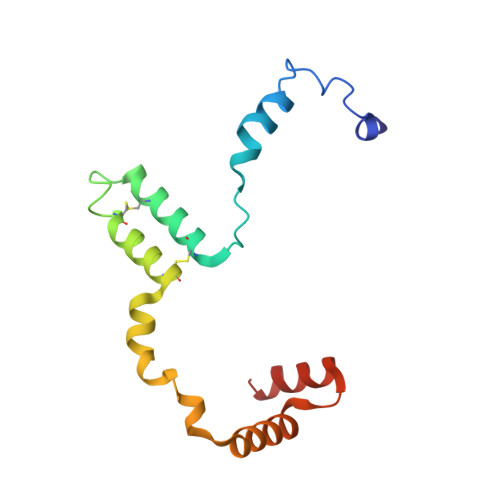
Euglena gracilis, a model organism of the eukaryotic supergroup Discoba harbouring also clinically important parasitic species, possesses diverse metabolic strategies and an atypical electron transport chain. While structures of the electron transport chain complexes and supercomplexes of most other eukaryotic clades have been reported, no similar structure is currently available for Discoba, limiting the understandings of its core metabolism and leaving a gap in the evolutionary tree of eukaryotic bioenergetics. Here, we report high-resolution cryo-EM structures of Euglena's respirasome I + III 2 + IV and supercomplex III 2 + IV 2 . A previously unreported fatty acid synthesis domain locates on the tip of complex I's peripheral arm, providing a clear picture of its atypical subunit composition identified previously. Individual complexes are re-arranged in the respirasome to adapt to the non-uniform membrane curvature of the discoidal cristae. Furthermore, Euglena's conformationally rigid complex I is deactivated by restricting ubiquinone's access to its substrate tunnel. Our findings provide structural insights for therapeutic developments against euglenozoan parasite infections.
- Department of Biophysics and Department of Critical Care Medicine of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, Hangzhou, 310058, China.
Organizational Affiliation: