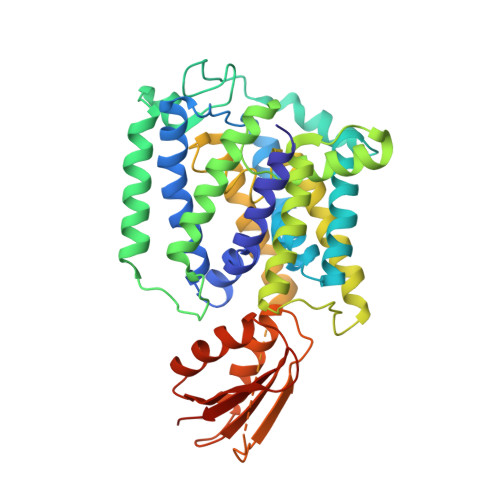
YeeD is an essential partner for YeeE-mediated thiosulfate uptake in bacteria and regulates thiosulfate ion decomposition.
Ikei, M., Miyazaki, R., Monden, K., Naito, Y., Takeuchi, A., Takahashi, Y.S., Tanaka, Y., Murata, K., Mori, T., Ichikawa, M., Tsukazaki, T.(2024) PLoS Biol 22: e3002601-e3002601
- PubMed: 38656967 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.3002601
- Primary Citation Related Structures:
8J4C, 8K1R - PubMed Abstract:
Uptake of thiosulfate ions as an inorganic sulfur source from the environment is important for bacterial sulfur assimilation. Recently, a selective thiosulfate uptake pathway involving a membrane protein YeeE (TsuA) in Escherichia coli was characterized. YeeE-like proteins are conserved in some bacteria, archaea, and eukaryotes. However, the precise function of YeeE, along with its potential partner protein in the thiosulfate ion uptake pathway, remained unclear. Here, we assessed selective thiosulfate transport via Spirochaeta thermophila YeeE in vitro and characterized E. coli YeeD (TsuB) as an adjacent and essential protein for YeeE-mediated thiosulfate uptake in vivo. We further showed that S. thermophila YeeD possesses thiosulfate decomposition activity and that a conserved cysteine in YeeD was modified to several forms in the presence of thiosulfate. Finally, the crystal structures of S. thermophila YeeE-YeeD fusion proteins at 3.34-Å and 2.60-Å resolutions revealed their interactions. The association was evaluated by a binding assay using purified S. thermophila YeeE and YeeD. Based on these results, a model of the sophisticated uptake of thiosulfate ions by YeeE and YeeD is proposed.
- Division of Biological Science, Graduate School of Science and Technology, Nara Institute of Science and Technology, Ikoma, Nara, Japan.
Organizational Affiliation: