Structure of multidrug resistance-associated protein 2 at 3.62 Angstroms resolution
Yun, C.H., Gao, H.M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ATP-binding cassette sub-family C member 2 | 1,589 | Homo sapiens | Mutation(s): 1 Gene Names: ABCC2, CMOAT, CMOAT1, CMRP, MRP2 EC: 7.6.2 (PDB Primary Data), 7.6.2.2 (PDB Primary Data), 7.6.2.3 (PDB Primary Data) Membrane Entity: Yes | 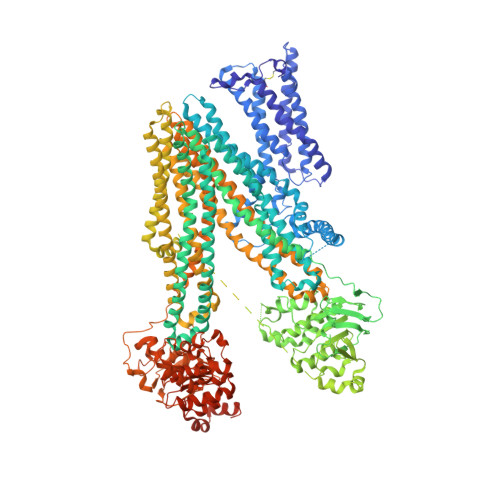 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q92887 GTEx: ENSG00000023839 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q92887 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | -- |