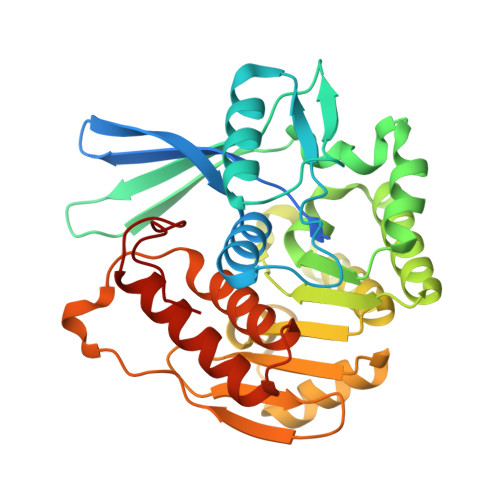
Structure of YdjH from Acinetobacter baumannii revealed an active site of YdjH family sugar kinase.
Lee, G.H., Kim, J.H., Ha, H.J., Park, H.H.(2023) Biochem Biophys Res Commun 664: 27-34
- PubMed: 37130458 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2023.04.073
- Primary Citation Related Structures:
8IWL - PubMed Abstract:
Bacterial sugar kinase is a central enzyme for proper sugar degradation in bacteria, essential for survival and growth. Therefore, this enzyme family is a primary target for antibacterial drug development, with YdjH most preferring to phosphorylate higher-order monosaccharides with a carboxylate terminus. Sugar kinases express diverse specificity and functions, making specificity determination of this family a prominent issue. This study examines the YdjH crystal structure from Acinetobacter baumannii (abYdjH), which has an exceptionally high antibiotic resistance and is considered a superbug. Our structural and biochemical study revealed that abYdjH has a widely open lid domain and is a solution dimer. In addition, the putative active site of abYdjH was determined based on structural analysis, sequence comparison, and in silico docking. Finally, we proposed the active site-forming residues that determine various sugar specificities from abYdjH. This study contributes towards a deeper understanding of the phosphorylation process and bacterial sugar metabolism of YdjH family to design the next-generation antibiotics for targeting A. baumannii.
- College of Pharmacy, Chung-Ang University, Seoul, 06974, Republic of Korea; Department of Global Innovative Drugs, Graduate School of Chung-Ang University, Seoul, 06974, Republic of Korea.
Organizational Affiliation: