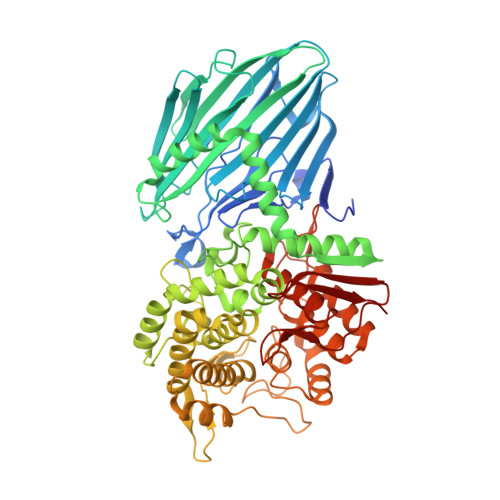
Bacteroidota polysaccharide utilization system for branched dextran exopolysaccharides from lactic acid bacteria.
Nakamura, S., Kurata, R., Tonozuka, T., Funane, K., Park, E.Y., Miyazaki, T.(2023) J Biological Chem 299: 104885-104885
- PubMed: 37269952 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.104885
- Primary Citation Related Structures:
8IU8, 8IU9, 8IUA, 8IUB, 8IUC - PubMed Abstract:
Dextran is an α-(1→6)-glucan that is synthesized by some lactic acid bacteria, and branched dextran with α-(1→2)-, α-(1→3)-, and α-(1→4)-linkages are often produced. Although many dextranases are known to act on the α-(1→6)-linkage of dextran, few studies have functionally analyzed the proteins involved in degrading branched dextran. The mechanism by which bacteria utilize branched dextran is unknown. Earlier, we identified dextranase (FjDex31A) and kojibiose hydrolase (FjGH65A) in the dextran utilization locus (FjDexUL) of a soil Bacteroidota Flavobacterium johnsoniae and hypothesized that FjDexUL is involved in the degradation of α-(1→2)-branched dextran. In this study, we demonstrate that FjDexUL proteins recognize and degrade α-(1→2)- and α-(1→3)-branched dextrans produced by Leuconostoc citreum S-32 (S-32 α-glucan). The FjDexUL genes were significantly upregulated when S-32 α-glucan was the carbon source compared with α-glucooligosaccharides and α-glucans, such as linear dextran and branched α-glucan from L. citreum S-64. FjDexUL glycoside hydrolases synergistically degraded S-32 α-glucan. The crystal structure of FjGH66 shows that some sugar-binding subsites can accommodate α-(1→2)- and α-(1→3)-branches. The structure of FjGH65A in complex with isomaltose supports that FjGH65A acts on α-(1→2)-glucosyl isomaltooligosaccharides. Furthermore, two cell surface sugar-binding proteins (FjDusD and FjDusE) were characterized, and FjDusD showed an affinity for isomaltooligosaccharides and FjDusE for dextran, including linear and branched dextrans. Collectively, FjDexUL proteins are suggested to be involved in the degradation of α-(1→2)- and α-(1→3)-branched dextrans. Our results will be helpful in understanding the bacterial nutrient requirements and symbiotic relationships between bacteria at the molecular level.
- Department of Bioscience, Graduate School of Science and Technology, Shizuoka University, Suruga-ku, Shizuoka, Japan.
Organizational Affiliation: