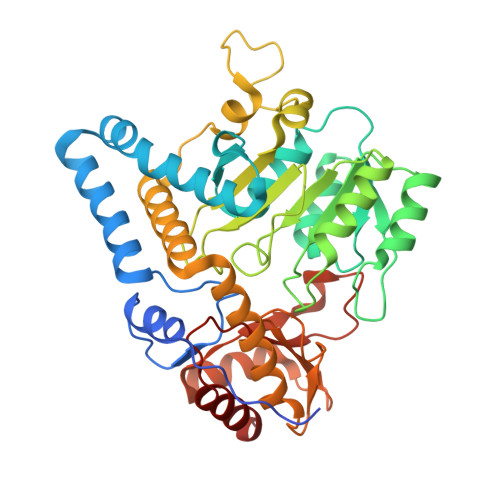
Structure of mycobacterial ergothioneine-biosynthesis C-S lyase EgtE.
Wei, L., Liu, L., Gong, W.(2024) J Biological Chem 300: 105539-105539
- PubMed: 38072054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.105539
- Primary Citation Related Structures:
8IRK, 8IRY, 8IRZ, 8IS0 - PubMed Abstract:
L-ergothioneine is widely distributed among various microbes to regulate their physiology and pathogenicity within complex environments. One of the key steps in the ergothioneine-biosynthesis pathway, the C-S bond cleavage reaction, uses the pyridoxal 5'-phosphate dependent C-S lyase to produce the final product L-ergothioneine. Here, we present the crystallographic structure of the ergothioneine-biosynthesis C-S lyase EgtE from Mycobacterium smegmatis (MsEgtE) represents the first published structure of ergothioneine-biosynthesis C-S lyases in bacteria and shows the effects of active site residues on the enzymatic reaction. The MsEgtE and the previously reported ergothioneine-biosynthesis C-S lyase Egt2 from Neurospora crassa (NcEgt2) fold similarly. However, discrepancies arise in terms of substrate recognition, as observed through sequence and structure comparison of MsEgtE and NcEgt2. The structural-based sequence alignment of the ergothioneine-biosynthesis C-S lyase from fungi and bacteria shows clear distinctions among the recognized substrate residues, but Arg348 is critical and an extremely conserved residue for substrate recognition. The α14 helix is exclusively found in the bacteria EgtE, which represent the most significant difference between bacteria EgtE and fungi Egt2, possibly resulting from the convergent evolution of bacteria and fungi.
- School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, China.
Organizational Affiliation: