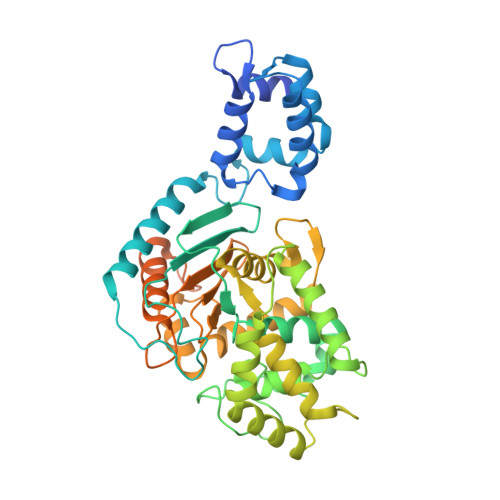
Structural basis for activation and filamentation of glutaminase.
Guo, C.J., Wang, Z.X., Liu, J.L.(2024) Cell Res 34: 76-79
- PubMed: 37833360 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-023-00886-0
- Primary Citation Related Structures:
8IMA, 8IMB - School of Life Science and Technology, ShanghaiTech University, Shanghai, China.
Organizational Affiliation: