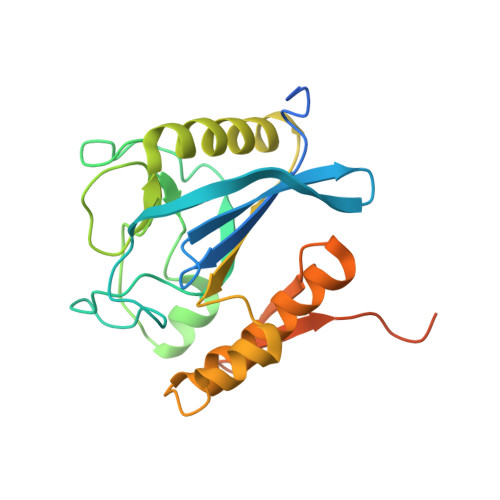
Structural basis for the hydrolytic activity of the transpeptidase-like protein DpaA to detach Braun's lipoprotein from peptidoglycan.
Wang, H.J., Hernandez-Rocamora, V.M., Kuo, C.I., Hsieh, K.Y., Lee, S.H., Ho, M.R., Tu, Z., Vollmer, W., Chang, C.I.(2023) mBio 14: e0137923-e0137923
- PubMed: 37830798 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.01379-23
- Primary Citation Related Structures:
8IKR - PubMed Abstract:
Cross-linking reaction of Braun's lipoprotein (Lpp) to peptidoglycan (PG) is catalyzed by some members of the YkuD family of transpeptidases. However, the exact opposite reaction of cleaving the Lpp-PG cross-link is performed by DpaA, which is also a YkuD-like protein. In this work, we determined the crystal structure of DpaA to provide the molecular rationale for the ability of the transpeptidase-like protein to cleave, rather than form, the Lpp-PG linkage. Our findings also revealed the structural features that distinguish the different functional types of the YkuD family enzymes from one another.
- Institute of Biological Chemistry, Academia Sinica , Taipei, Taiwan.
Organizational Affiliation: