Crystal structure of protease-associated domain of Arabidopsis vacuolar sorting receptor 1 at pH 4.6
Tsao, H.E., Lui, S.N., Wong, K.B.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Vacuolar-sorting receptor 1 | 165 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: VSR1, BP80B, ELP, ELP1, At3g52850, F8J2.20 | 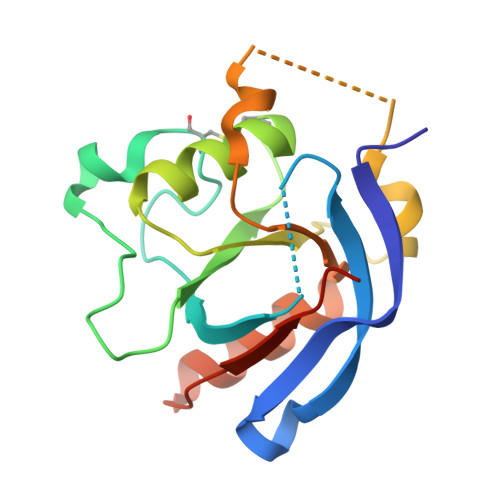 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P93026 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EPE Download:Ideal Coordinates CCD File | E [auth A] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | C [auth A], F [auth A], G [auth A], H [auth A], L [auth A] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | B [auth A], I [auth A], J [auth A], K [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| CL Download:Ideal Coordinates CCD File | D [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 32.987 | α = 90 |
| b = 60.318 | β = 103.36 |
| c = 38.466 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| The University Grants Committee, Research Grants Council (RGC) | Hong Kong | CUHK14151416 |
| The University Grants Committee, Research Grants Council (RGC) | Hong Kong | C4041-18EF |
| The University Grants Committee, Research Grants Council (RGC) | Hong Kong | AoE/M-05/12 |
| The University Grants Committee, Research Grants Council (RGC) | Hong Kong | AoE/M403/16 |
| The University Grants Committee, Research Grants Council (RGC) | Hong Kong | C4033-19E |