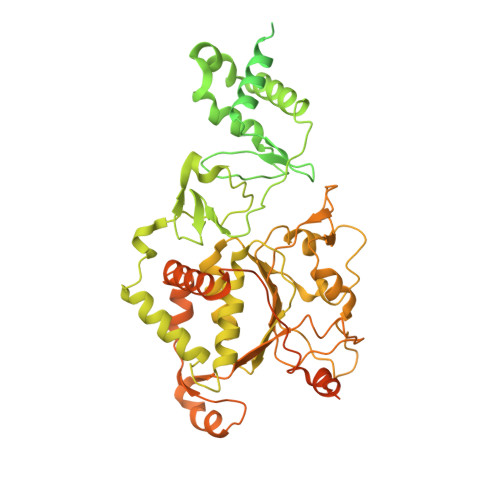
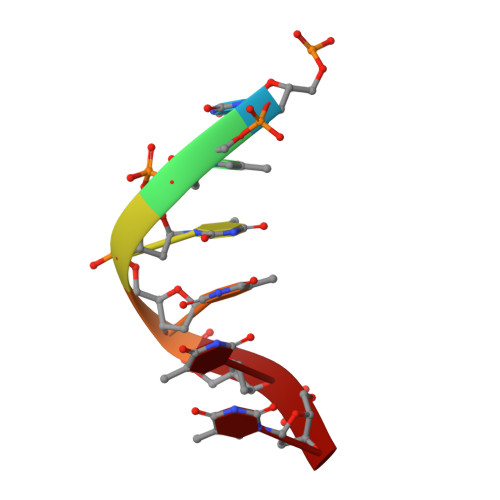
Structural insight into the assembly and working mechanism of helicase-primase D5 from Mpox virus.
Li, Y., Zhu, J., Guo, Y., Yan, R.(2024) Nat Struct Mol Biol 31: 68-81
- PubMed: 38177671 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01142-0
- Primary Citation Related Structures:
8HWA, 8HWB, 8HWC, 8HWD, 8HWE, 8HWF, 8HWG, 8HWH - PubMed Abstract:
The Mpox pandemic, caused by the Mpox virus (or monkeypox virus, MPXV), has gained global attention. The D5 protein, a putative helicase-primase found in MPXV, plays a vital role in viral replication and genome uncoating. Here we determined multiple cryo-EM structures of full-length hexameric D5 in diverse states. These states were captured during ATP hydrolysis while moving along the single-stranded DNA (ssDNA) track. Through comprehensive structural analysis combined with the helicase activity system, we revealed that when the primase domain is truncated or the interaction between the primase and helicase domains is disrupted, the double-stranded DNA (dsDNA) unwinds into ssDNA, suggesting a critical regulatory role of the primase domain. Two transition states bound with ssDNA substrate during unwinding reveals that two ATP molecules were consumed to drive DNA moving forward two nucleotides. Collectively, our findings shed light on the molecular mechanism that links ATP hydrolysis to the DNA unwinding in poxviruses.
- Department of Biochemistry, School of Medicine, Key University Laboratory of Metabolism and Health of Guangdong, Institute for Biological Electron Microscopy, Southern University of Science and Technology, Shenzhen, China.
Organizational Affiliation: