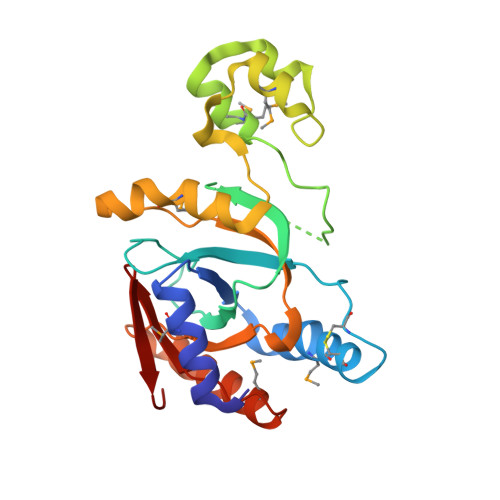
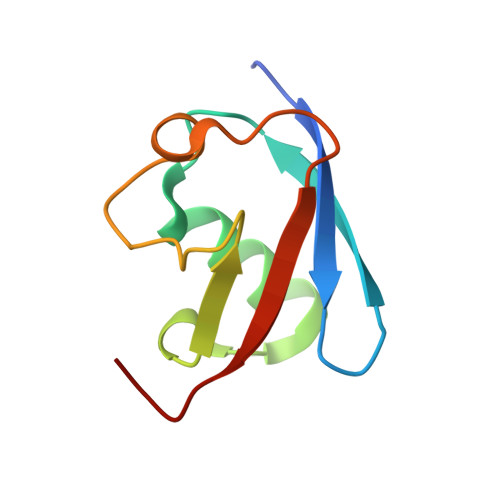
Molecular basis of threonine ADP-ribosylation of ubiquitin by bacterial ARTs.
Tan, J., Xu, Y., Wang, X., Yan, F., Xian, W., Liu, X., Chen, Y., Zhu, Y., Zhou, Y.(2024) Nat Chem Biol 20: 463-472
- PubMed: 37945894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01475-3
- Primary Citation Related Structures:
8HTC, 8HTD, 8HTE, 8HTF - PubMed Abstract:
Ubiquitination plays essential roles in eukaryotic cellular processes. The effector protein CteC from Chromobacterium violaceum blocks host ubiquitination by mono-ADP-ribosylation of ubiquitin (Ub) at residue T66. However, the structural basis for this modification is unknown. Here we report three crystal structures of CteC in complexes with Ub, NAD + or ADP-ribosylated Ub, which represent different catalytic states of CteC in the modification. CteC adopts a special 'D-E' catalytic motif for catalysis and binds NAD + in a half-ligand binding mode. The specific recognition of Ub by CteC is determined by a relatively separate Ub-targeting domain and a long loop L6, not the classic ADP-ribosylating turn-turn loop. Structural analyses with biochemical results reveal that CteC represents a large family of poly (ADP-ribose) polymerase (PARP)-like ADP-ribosyltransferases, which harbors chimeric features from the R-S-E and H-Y-E classes of ADP-ribosyltransferases. The family of CteC-like ADP-ribosyltransferases has a common 'D-E' catalytic consensus and exists extensively in bacteria and eukaryotic microorganisms.
- The MOE Key Laboratory for Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, Life Sciences Institute, Zhejiang University, Hangzhou, China.
Organizational Affiliation: