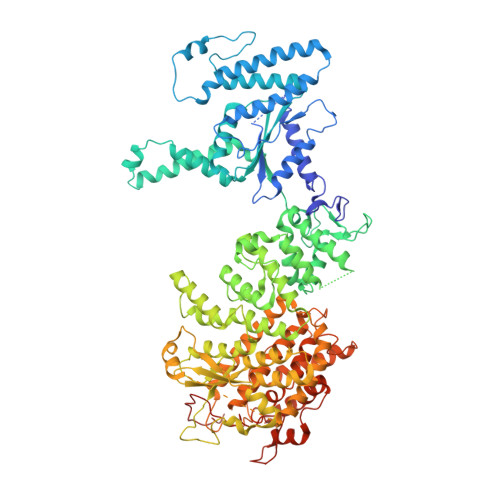
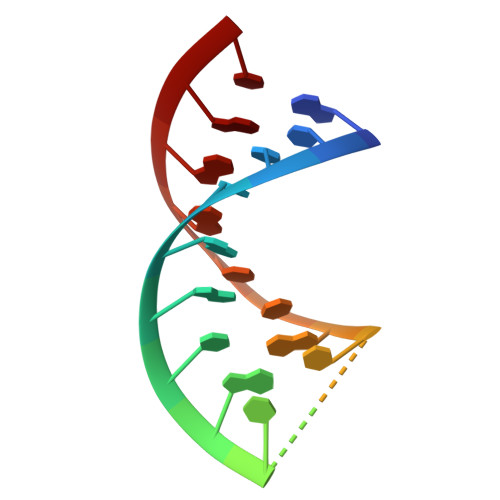
Molecular basis of RADAR anti-phage supramolecular assemblies.
Gao, Y., Luo, X., Li, P., Li, Z., Ye, F., Liu, S., Gao, P.(2023) Cell 186: 999-1012.e20
- PubMed: 36764292 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2023.01.026
- Primary Citation Related Structures:
8HR7, 8HR8, 8HR9, 8HRA, 8HRB, 8HRC - PubMed Abstract:
Adenosine-to-inosine RNA editing has been proposed to be involved in a bacterial anti-phage defense system called RADAR. RADAR contains an adenosine triphosphatase (RdrA) and an adenosine deaminase (RdrB). Here, we report cryo-EM structures of RdrA, RdrB, and currently identified RdrA-RdrB complexes in the presence or absence of RNA and ATP. RdrB assembles into a dodecameric cage with catalytic pockets facing outward, while RdrA adopts both autoinhibited tetradecameric and activation-competent heptameric rings. Structural and functional data suggest a model in which RNA is loaded through the bottom section of the RdrA ring and translocated along its inner channel, a process likely coupled with ATP-binding status. Intriguingly, up to twelve RdrA rings can dock one RdrB cage with precise alignments between deaminase catalytic pockets and RNA-translocation channels, indicative of enzymatic coupling of RNA translocation and deamination. Our data uncover an interesting mechanism of enzymatic coupling and anti-phage defense through supramolecular assemblies.
- CAS Key Laboratory of Infection and Immunity, National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: