Structural basis for the assembly of the DNA polymerase holoenzyme from a monkeypox virus variant.
Li, Y., Shen, Y., Hu, Z., Yan, R.(2023) Sci Adv 9: eadg2331-eadg2331
- PubMed: 37075110 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adg2331
- Primary Citation Related Structures:
8HLZ, 8HM0 - PubMed Abstract:
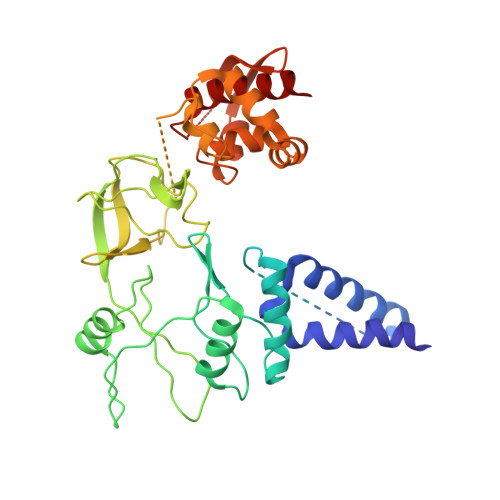
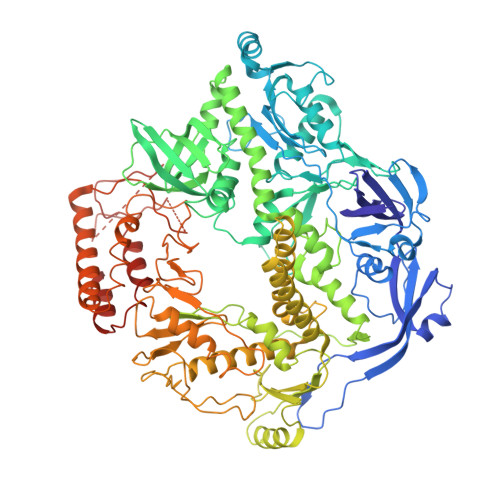
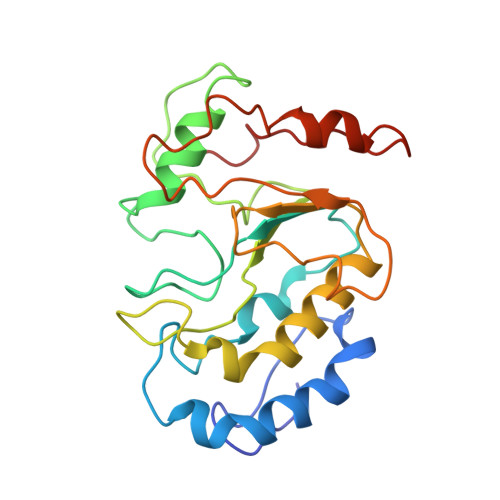
The ongoing global pandemic caused by a variant of the monkeypox (or mpox) virus (MPXV) has prompted widespread concern. The MPXV DNA polymerase holoenzyme, consisting of F8, A22, and E4, is vital for replicating the viral genome and represents a crucial target for the development of antiviral drugs. However, the assembly and working mechanism for the DNA polymerase holoenzyme of MPXV remains elusive. Here, we present the cryo-electron microscopy (cryo-EM) structure of the DNA polymerase holoenzyme at an overall resolution of 3.5 Å. Unexpectedly, the holoenzyme is assembled as a dimer of heterotrimers, of which the extra interface between the thumb domain of F8 and A22 shows a clash between A22 and substrate DNA, suggesting an autoinhibition state. Addition of exogenous double-stranded DNA shifts the hexamer into trimer exposing DNA binding sites, potentially representing a more active state. Our findings provide crucial steps toward developing targeted antiviral therapies for MPXV and related viruses.
- Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou, Zhejiang Province 310024, China.
Organizational Affiliation: