Structural insight into the septal peptidoglycan hydrolysis machinery of bacterial cell division
Zhang, Z., Dong, H., Chen, Y.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cell division ATP-binding protein FtsE | 222 | Escherichia coli K-12 | Mutation(s): 1 Gene Names: ftsE, b3463, JW3428 Membrane Entity: Yes | 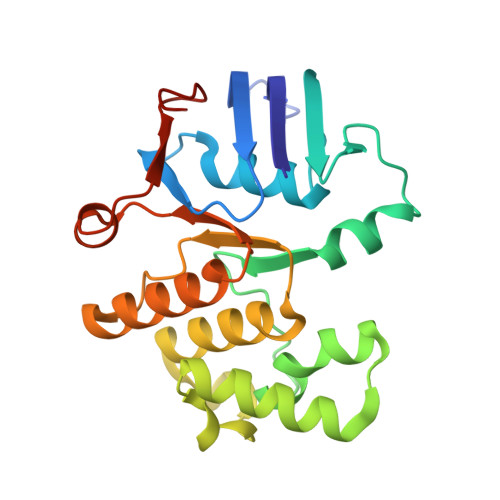 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A9R7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cell division protein FtsX | 352 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: ftsX, ftsS, b3462, JW3427 Membrane Entity: Yes | 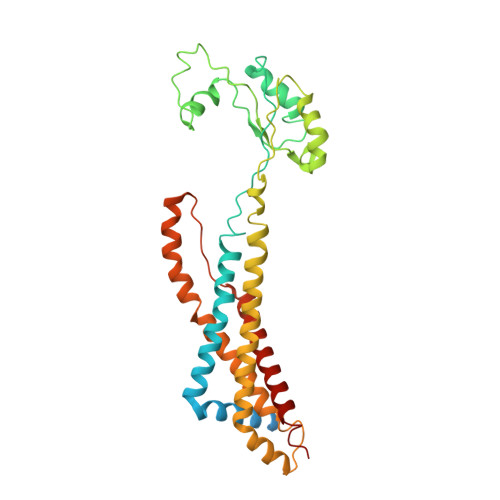 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0AC30 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Murein hydrolase activator EnvC | 419 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: envC, yibP, b3613, JW5646 Membrane Entity: Yes | 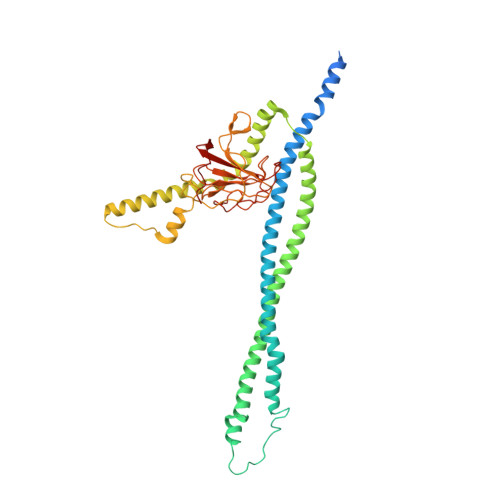 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P37690 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth A], G [auth B] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 31800052 |
| Ministry of Science and Technology (MoST, China) | China | 2018YFA093200 |