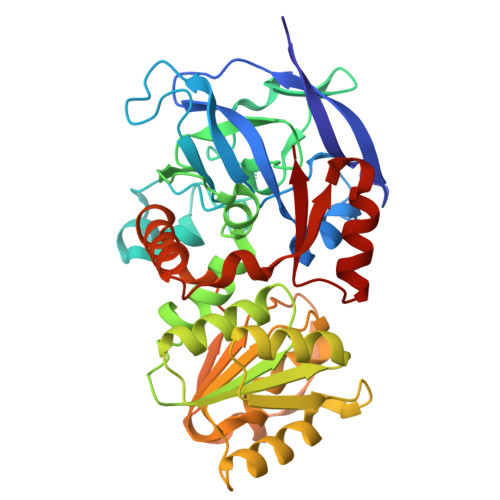
Unique alcohol dehydrogenases involved in algal sugar utilization by marine bacteria
Brott, S., Nam, K.H., Thomas, F., Dutschei, T., Reisky, L., Behrens, M., Grimm, H.C., Michel, G., Schweder, T., Bornscheuer, U.T.(2023) Appl Microbiol Biotechnol 107: 2363-2384