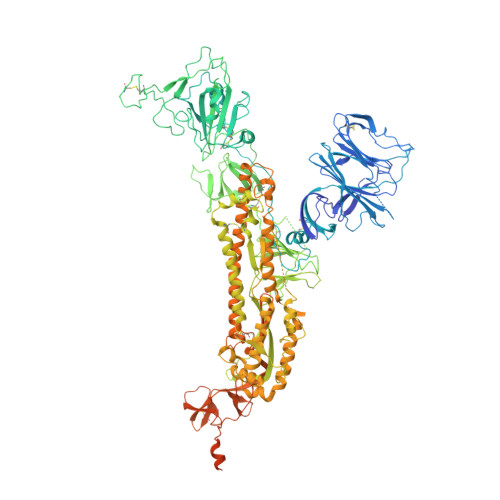
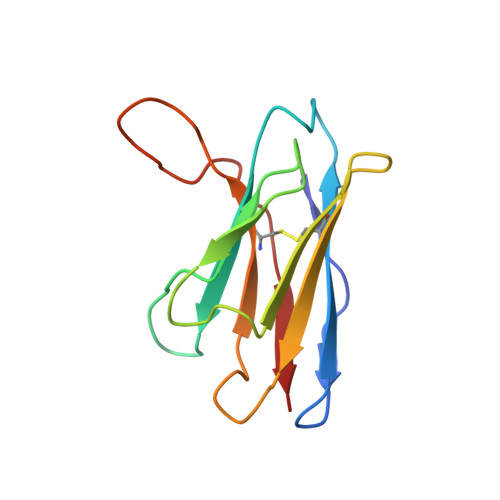
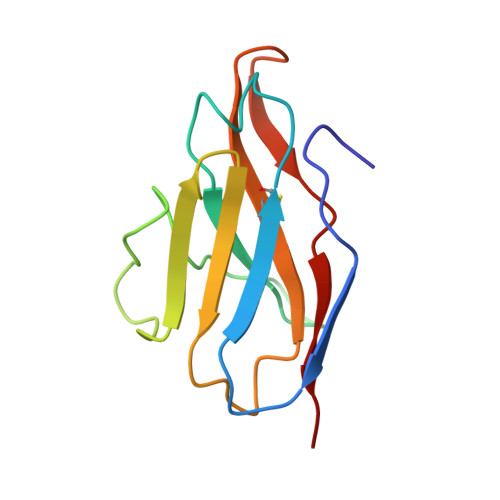
Comprehensive structural analysis reveals broad-spectrum neutralizing antibodies against SARS-CoV-2 Omicron variants.
Chi, X., Xia, L., Zhang, G., Chi, X., Huang, B., Zhang, Y., Chen, Z., Han, J., Wu, L., Li, Z., Sun, H., Huang, P., Yu, C., Chen, W., Zhou, Q.(2023) Cell Discov 9: 37-37
- PubMed: 37015915 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-023-00535-1
- Primary Citation Related Structures:
8GTO, 8GTP, 8GTQ - PubMed Abstract:
The pandemic of COVID-19 caused by SARS-CoV-2 continues to spread around the world. Mutant strains of SARS-CoV-2 are constantly emerging. At present, Omicron variants have become mainstream. In this work, we carried out a systematic and comprehensive analysis of the reported spike protein antibodies, counting the epitopes and genotypes of these antibodies. We further comprehensively analyzed the impact of Omicron mutations on antibody epitopes and classified these antibodies according to their binding patterns. We found that the epitopes of the H-RBD class antibodies were significantly less affected by Omicron mutations than other classes. Binding and virus neutralization experiments showed that such antibodies could effectively inhibit the immune escape of Omicron. Cryo-EM results showed that this class of antibodies utilized a conserved mechanism to neutralize SARS-CoV-2. Our results greatly help us deeply understand the impact of Omicron mutations. Meanwhile, it also provides guidance and insights for developing Omicron antibodies and vaccines.
- Institute of Biotechnology, Academy of Military Medical Sciences, Beijing, China.
Organizational Affiliation: