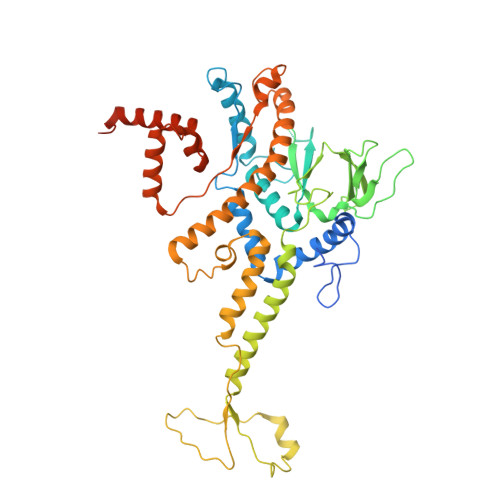
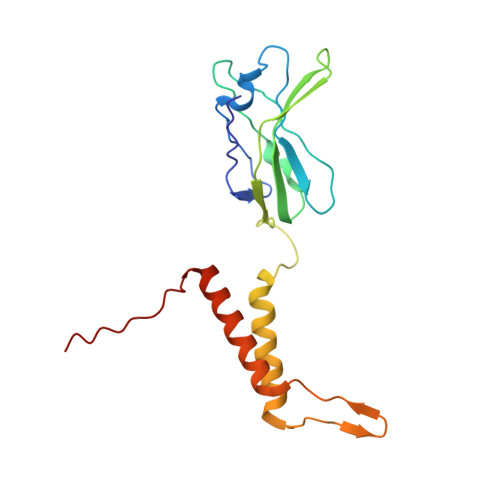
Structure and proposed DNA delivery mechanism of a marine roseophage.
Huang, Y., Sun, H., Wei, S., Cai, L., Liu, L., Jiang, Y., Xin, J., Chen, Z., Que, Y., Kong, Z., Li, T., Yu, H., Zhang, J., Gu, Y., Zheng, Q., Li, S., Zhang, R., Xia, N.(2023) Nat Commun 14: 3609-3609
- PubMed: 37330604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39220-y
- Primary Citation Related Structures:
8GTA, 8GTB, 8GTC, 8GTD, 8GTF - PubMed Abstract:
Tailed bacteriophages (order, Caudovirales) account for the majority of all phages. However, the long flexible tail of siphophages hinders comprehensive investigation of the mechanism of viral gene delivery. Here, we report the atomic capsid and in-situ structures of the tail machine of the marine siphophage, vB_DshS-R4C (R4C), which infects Roseobacter. The R4C virion, comprising 12 distinct structural protein components, has a unique five-fold vertex of the icosahedral capsid that allows genome delivery. The specific position and interaction pattern of the tail tube proteins determine the atypical long rigid tail of R4C, and further provide negative charge distribution within the tail tube. A ratchet mechanism assists in DNA transmission, which is initiated by an absorption device that structurally resembles the phage-like particle, RcGTA. Overall, these results provide in-depth knowledge into the intact structure and underlining DNA delivery mechanism for the ecologically important siphophages.
- State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics, School of Public Health, School of Life Sciences, Xiamen University, Xiamen, 361102, China.
Organizational Affiliation: