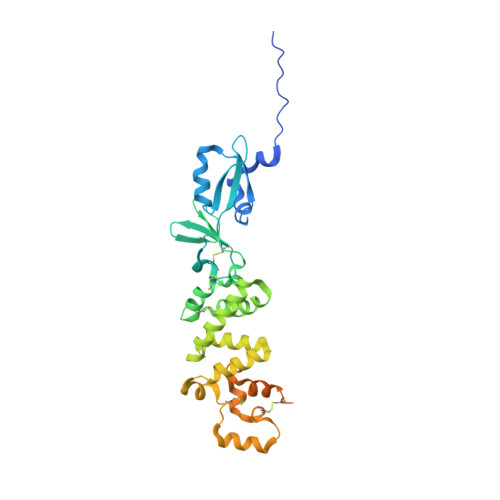
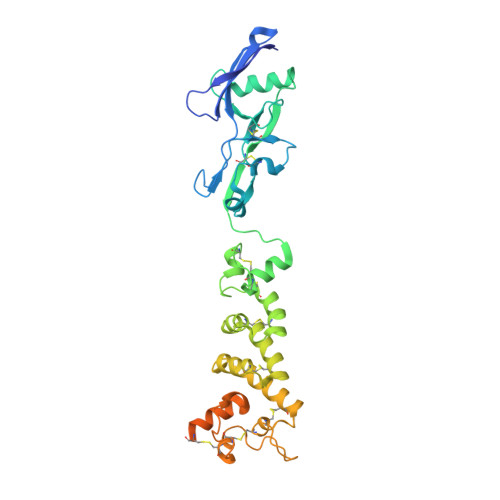
Structural basis of poxvirus A16/G9 binding for sub-complex formation.
Yang, F., Lin, S., Chen, Z., Yue, D., Yang, M., He, B., Cao, Y., Dong, H., Li, J., Zhao, Q., Lu, G.(2023) Emerg Microbes Infect 12: 2179351-2179351
- PubMed: 36757688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/22221751.2023.2179351
- Primary Citation Related Structures:
8GP6 - West China Hospital Emergency Department (WCHED), State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, People's Republic of China.
Organizational Affiliation: