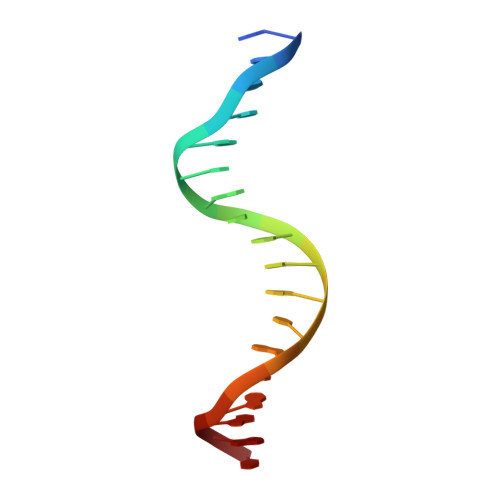
The structural basis for 2'-5'/3'-5'-cGAMP synthesis by cGAS.
Wu, S., Gabelli, S.B., Sohn, J.(2024) Nat Commun 15: 4012-4012
- PubMed: 38740774 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-48365-3
- Primary Citation Related Structures:
8EAE, 8G10, 8G1J, 8G23, 8GIM, 8GIN, 8GIO, 8GIP, 8GIR, 8GIS, 8GIT, 8SHK, 8SHU, 8SHY, 8SHZ, 8SI0, 8SJ0, 8SJ1, 8SJ2, 8SJ8, 8SKT - PubMed Abstract:
cGAS activates innate immune responses against cytosolic double-stranded DNA. Here, by determining crystal structures of cGAS at various reaction stages, we report a unifying catalytic mechanism. apo-cGAS assumes an array of inactive conformations and binds NTPs nonproductively. Dimerization-coupled double-stranded DNA-binding then affixes the active site into a rigid lock for productive metal•substrate binding. A web-like network of protein•NTP, intra-NTP, and inter-NTP interactions ensures the stepwise synthesis of 2'-5'/3'-5'-linked cGAMP while discriminating against noncognate NTPs and off-pathway intermediates. One divalent metal is sufficient for productive substrate binding, and capturing the second divalent metal is tightly coupled to nucleotide and linkage specificities, a process which manganese is preferred over magnesium by 100-fold. Additionally, we elucidate how mouse cGAS achieves more stringent NTP and linkage specificities than human cGAS. Together, our results reveal that an adaptable, yet precise lock-and-key-like mechanism underpins cGAS catalysis.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Organizational Affiliation: