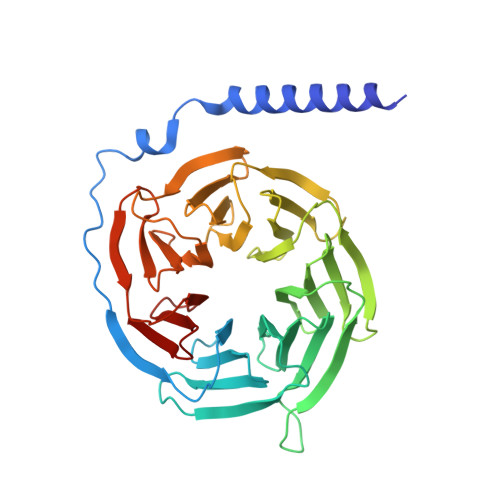
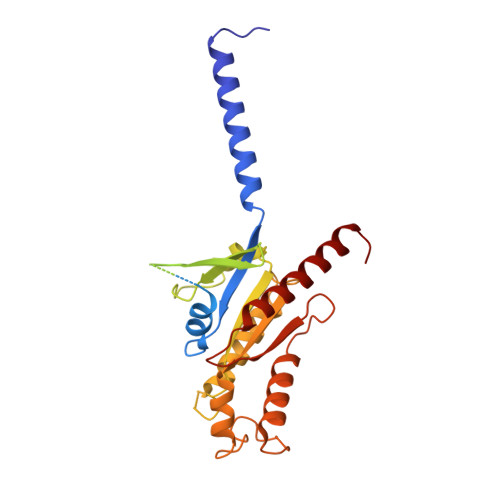
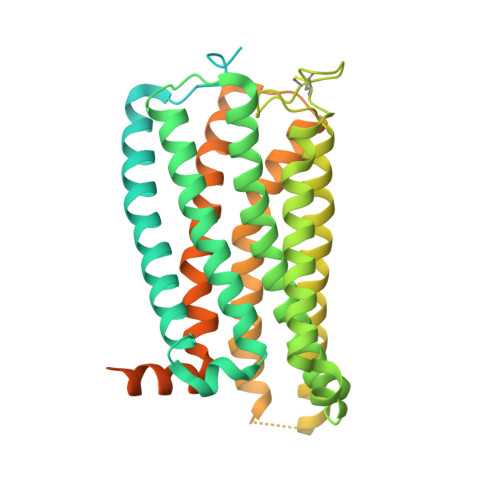
Structural basis for activation of CB1 by an endocannabinoid analog.
Krishna Kumar, K., Robertson, M.J., Thadhani, E., Wang, H., Suomivuori, C.M., Powers, A.S., Ji, L., Nikas, S.P., Dror, R.O., Inoue, A., Makriyannis, A., Skiniotis, G., Kobilka, B.(2023) Nat Commun 14: 2672-2672
- PubMed: 37160876 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-37864-4
- Primary Citation Related Structures:
8GHV - PubMed Abstract:
Endocannabinoids (eCBs) are endogenous ligands of the cannabinoid receptor 1 (CB1), a G protein-coupled receptor that regulates a number of therapeutically relevant physiological responses. Hence, understanding the structural and functional consequences of eCB-CB1 interactions has important implications for designing effective drugs targeting this receptor. To characterize the molecular details of eCB interaction with CB1, we utilized AMG315, an analog of the eCB anandamide to determine the structure of the AMG315-bound CB1 signaling complex. Compared to previous structures, the ligand binding pocket shows some differences. Using docking, molecular dynamics simulations, and signaling assays we investigated the functional consequences of ligand interactions with the "toggle switch" residues F200 3.36 and W356 6.48 . Further, we show that ligand-TM2 interactions drive changes to residues on the intracellular side of TM2 and are a determinant of efficacy in activating G protein. These intracellular TM2 rearrangements are unique to CB1 and are exploited by a CB1-specific allosteric modulator.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, 279 Campus Drive, Stanford, CA, 94305, USA.
Organizational Affiliation: