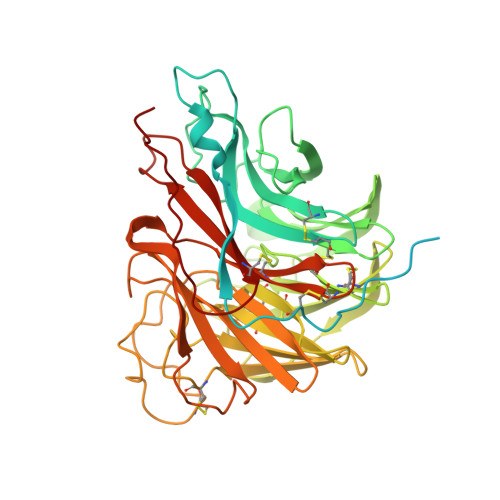
A pan-influenza antibody inhibiting neuraminidase via receptor mimicry.
Momont, C., Dang, H.V., Zatta, F., Hauser, K., Wang, C., di Iulio, J., Minola, A., Czudnochowski, N., De Marco, A., Branch, K., Donermeyer, D., Vyas, S., Chen, A., Ferri, E., Guarino, B., Powell, A.E., Spreafico, R., Yim, S.S., Balce, D.R., Bartha, I., Meury, M., Croll, T.I., Belnap, D.M., Schmid, M.A., Schaiff, W.T., Miller, J.L., Cameroni, E., Telenti, A., Virgin, H.W., Rosen, L.E., Purcell, L.A., Lanzavecchia, A., Snell, G., Corti, D., Pizzuto, M.S.(2023) Nature 618: 590-597
- PubMed: 37258672 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-06136-y
- Primary Citation Related Structures:
8G30, 8G3M, 8G3N, 8G3O, 8G3P, 8G3Q, 8G3R, 8G3V, 8G3Z, 8G40 - PubMed Abstract:
Rapidly evolving influenza A viruses (IAVs) and influenza B viruses (IBVs) are major causes of recurrent lower respiratory tract infections. Current influenza vaccines elicit antibodies predominantly to the highly variable head region of haemagglutinin and their effectiveness is limited by viral drift 1 and suboptimal immune responses 2 . Here we describe a neuraminidase-targeting monoclonal antibody, FNI9, that potently inhibits the enzymatic activity of all group 1 and group 2 IAVs, as well as Victoria/2/87-like, Yamagata/16/88-like and ancestral IBVs. FNI9 broadly neutralizes seasonal IAVs and IBVs, including the immune-evading H3N2 strains bearing an N-glycan at position 245, and shows synergistic activity when combined with anti-haemagglutinin stem-directed antibodies. Structural analysis reveals that D107 in the FNI9 heavy chain complementarity-determinant region 3 mimics the interaction of the sialic acid carboxyl group with the three highly conserved arginine residues (R118, R292 and R371) of the neuraminidase catalytic site. FNI9 demonstrates potent prophylactic activity against lethal IAV and IBV infections in mice. The unprecedented breadth and potency of the FNI9 monoclonal antibody supports its development for the prevention of influenza illness by seasonal and pandemic viruses.
- Vir Biotechnology, San Francisco, CA, USA.
Organizational Affiliation: