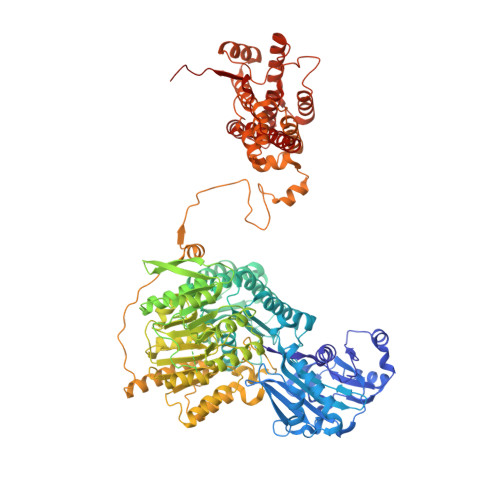
Allosteric role of the citrate synthase homology domain of ATP citrate lyase.
Wei, X., Schultz, K., Pepper, H.L., Megill, E., Vogt, A., Snyder, N.W., Marmorstein, R.(2023) Nat Commun 14: 2247-2247
- PubMed: 37076498 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-37986-9
- Primary Citation Related Structures:
7RIG, 7RKZ, 7RMP, 8G1E, 8G1F, 8G5C, 8G5D - PubMed Abstract:
ATP citrate lyase (ACLY) is the predominant nucleocytosolic source of acetyl-CoA and is aberrantly regulated in many diseases making it an attractive therapeutic target. Structural studies of ACLY reveal a central homotetrameric core citrate synthase homology (CSH) module flanked by acyl-CoA synthetase homology (ASH) domains, with ATP and citrate binding the ASH domain and CoA binding the ASH-CSH interface to produce acetyl-CoA and oxaloacetate products. The specific catalytic role of the CSH module and an essential D1026A residue contained within it has been a matter of debate. Here, we report biochemical and structural analysis of an ACLY-D1026A mutant demonstrating that this mutant traps a (3S)-citryl-CoA intermediate in the ASH domain in a configuration that is incompatible with the formation of acetyl-CoA, is able to convert acetyl-CoA and OAA to (3S)-citryl-CoA in the ASH domain, and can load CoA and unload acetyl-CoA in the CSH module. Together, this data support an allosteric role for the CSH module in ACLY catalysis.
- Department of Biochemistry & Biophysics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, 19104, USA.
Organizational Affiliation: