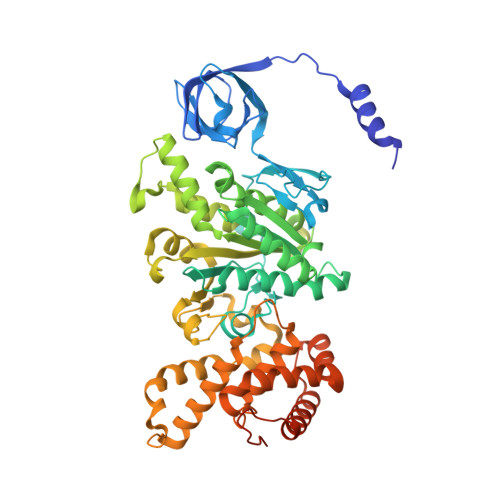
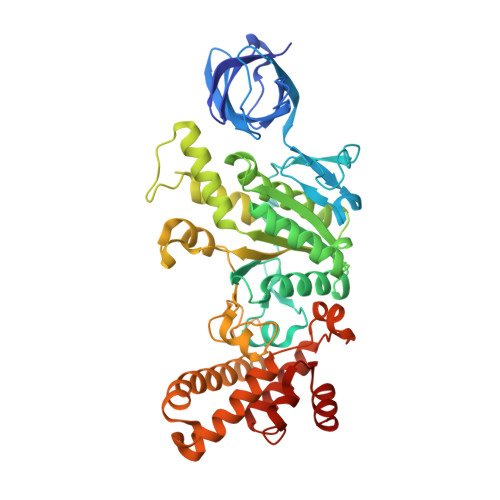
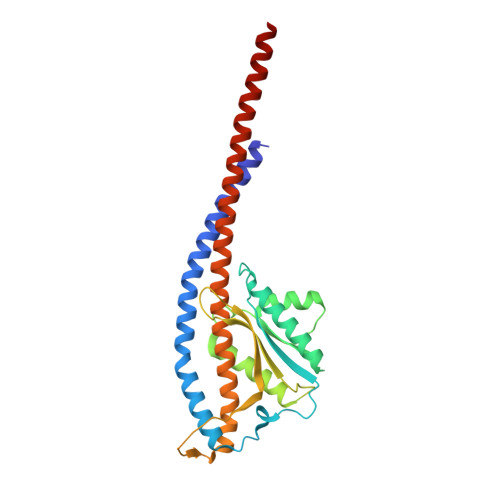
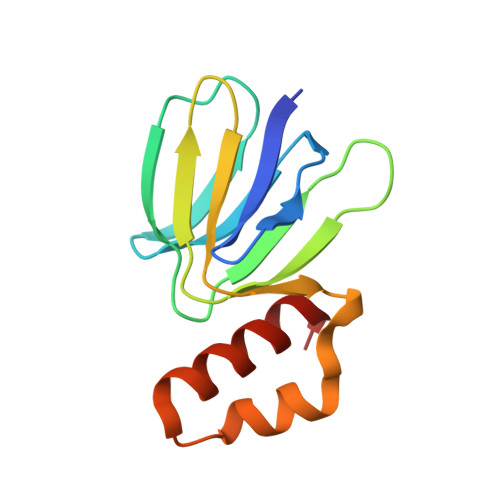
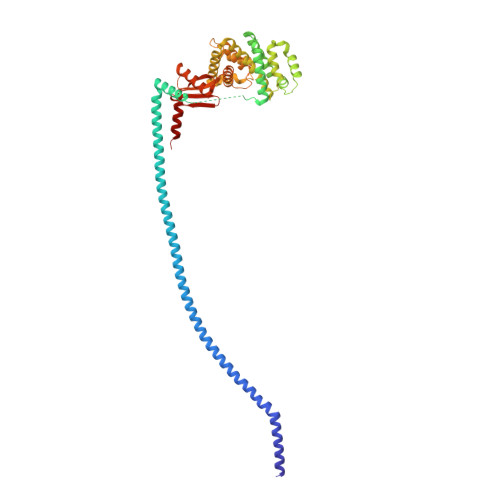
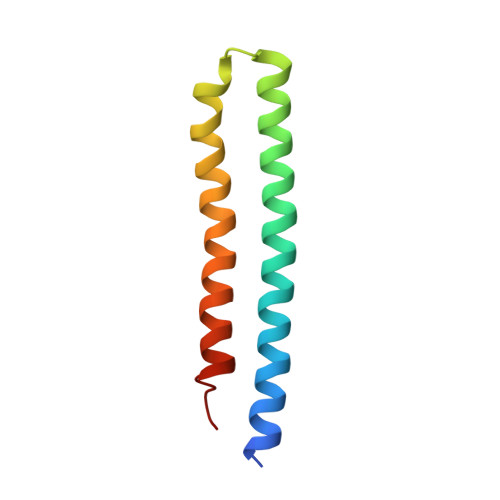
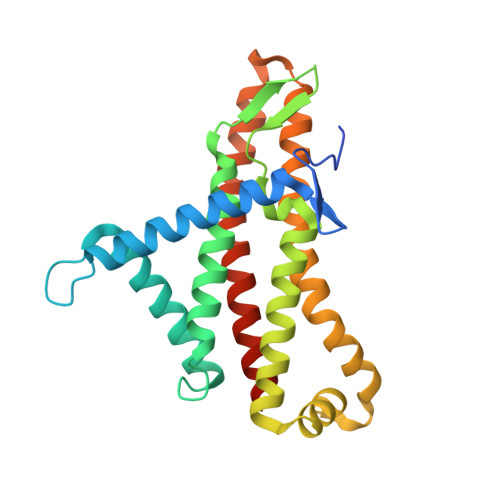
Mechanism of mycobacterial ATP synthase inhibition by squaramides and second generation diarylquinolines.
Courbon, G.M., Palme, P.R., Mann, L., Richter, A., Imming, P., Rubinstein, J.L.(2023) EMBO J 42: e113687-e113687
- PubMed: 37377118 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2023113687
- Primary Citation Related Structures:
8G07, 8G08, 8G09, 8G0A, 8G0B, 8G0C, 8G0D, 8G0E - PubMed Abstract:
Mycobacteria, such as Mycobacterium tuberculosis, depend on the activity of adenosine triphosphate (ATP) synthase for growth. The diarylquinoline bedaquiline (BDQ), a mycobacterial ATP synthase inhibitor, is an important medication for treatment of drug-resistant tuberculosis but suffers from off-target effects and is susceptible to resistance mutations. Consequently, both new and improved mycobacterial ATP synthase inhibitors are needed. We used electron cryomicroscopy and biochemical assays to study the interaction of Mycobacterium smegmatis ATP synthase with the second generation diarylquinoline TBAJ-876 and the squaramide inhibitor SQ31f. The aryl groups of TBAJ-876 improve binding compared with BDQ, while SQ31f, which blocks ATP synthesis ~10 times more potently than ATP hydrolysis, binds a previously unknown site in the enzyme's proton-conducting channel. Remarkably, BDQ, TBAJ-876, and SQ31f all induce similar conformational changes in ATP synthase, suggesting that the resulting conformation is particularly suited for drug binding. Further, high concentrations of the diarylquinolines uncouple the transmembrane proton motive force while for SQ31f they do not, which may explain why high concentrations of diarylquinolines, but not SQ31f, have been reported to kill mycobacteria.
- Molecular Medicine Program, The Hospital for Sick Children, Toronto, ON, Canada.
Organizational Affiliation: