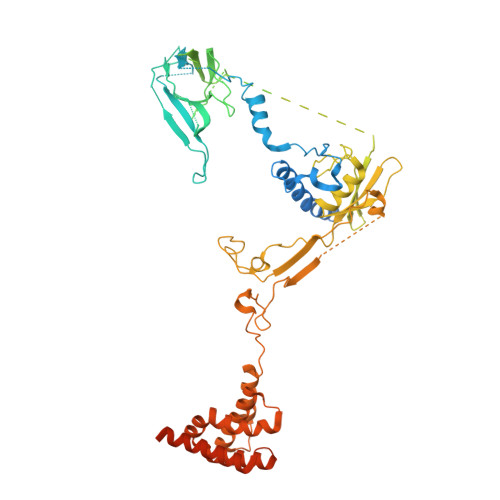
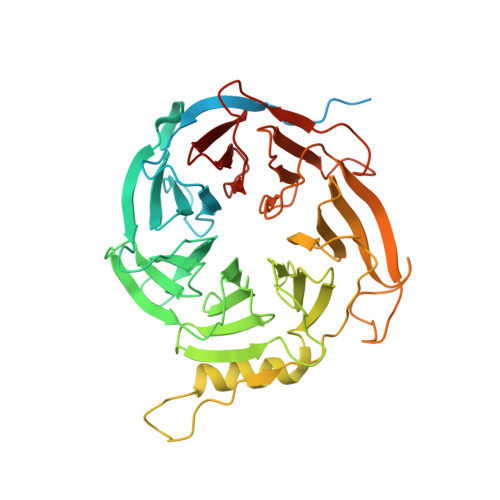
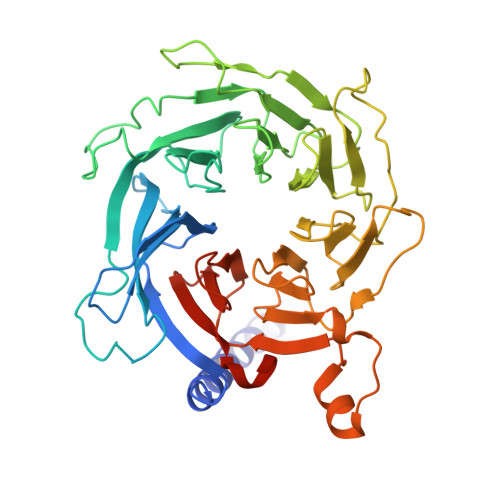
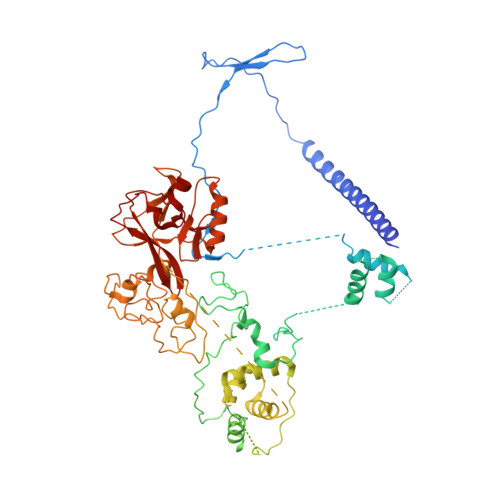
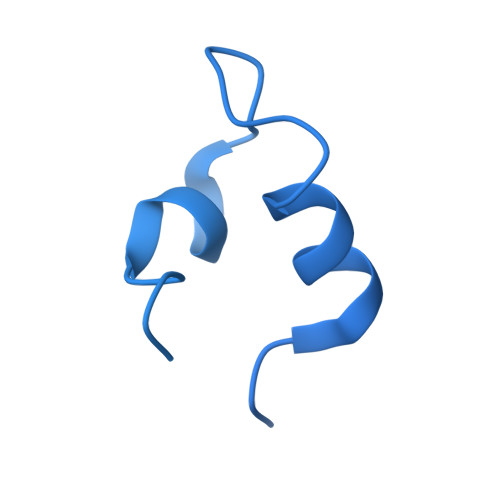
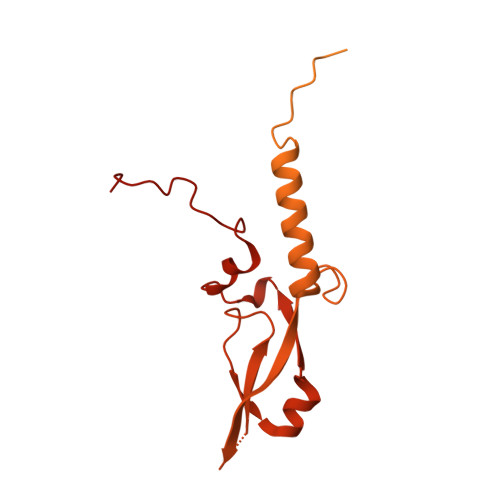
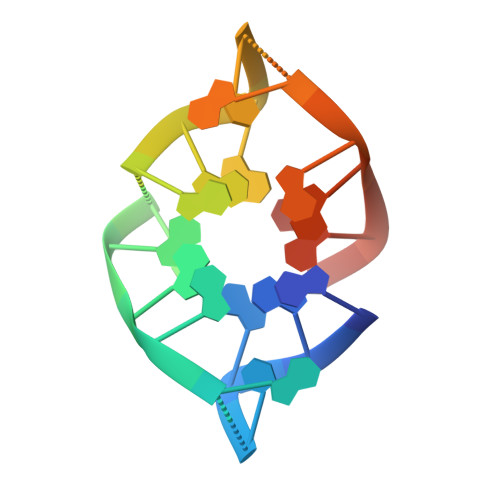Structural basis for inactivation of PRC2 by G-quadruplex RNA.
Song, J., Gooding, A.R., Hemphill, W.O., Love, B.D., Robertson, A., Yao, L., Zon, L.I., North, T.E., Kasinath, V., Cech, T.R.(2023) Science 381: 1331-1337
- PubMed: 37733873 Search on PubMed
- DOI: https://doi.org/10.1126/science.adh0059
- Primary Citation Related Structures:
8FYH - PubMed Abstract:
Polycomb repressive complex 2 (PRC2) silences genes through trimethylation of histone H3K27. PRC2 associates with numerous precursor messenger RNAs (pre-mRNAs) and long noncoding RNAs (lncRNAs) with a binding preference for G-quadruplex RNA. In this work, we present a 3.3-Å-resolution cryo-electron microscopy structure of PRC2 bound to a G-quadruplex RNA. Notably, RNA mediates the dimerization of PRC2 by binding both protomers and inducing a protein interface composed of two copies of the catalytic subunit EZH2, thereby blocking nucleosome DNA interaction and histone H3 tail accessibility. Furthermore, an RNA-binding loop of EZH2 facilitates the handoff between RNA and DNA, another activity implicated in PRC2 regulation by RNA. We identified a gain-of-function mutation in this loop that activates PRC2 in zebrafish. Our results reveal mechanisms for RNA-mediated regulation of a chromatin-modifying enzyme.
- Department of Biochemistry, University of Colorado Boulder, Boulder, CO 80303, USA.
Organizational Affiliation: