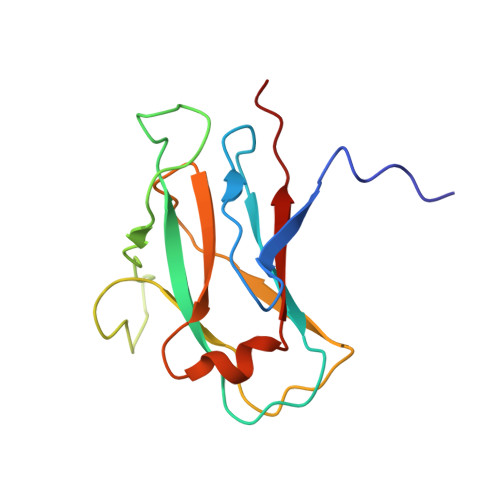
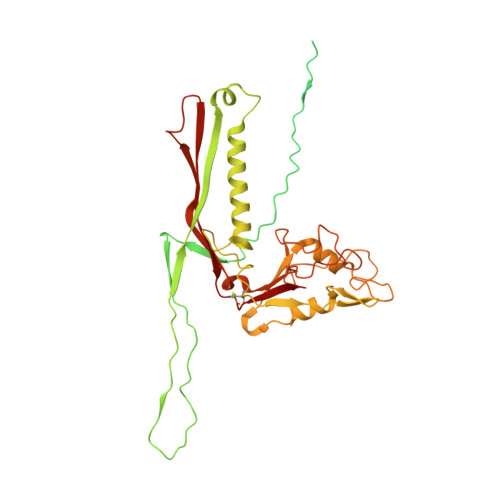
Neck and capsid architecture of the robust Agrobacterium phage Milano.
Sonani, R.R., Esteves, N.C., Horton, A.A., Kelly, R.J., Sebastian, A.L., Wang, F., Kreutzberger, M.A.B., Leiman, P.G., Scharf, B.E., Egelman, E.H.(2023) Commun Biol 6: 921-921
- PubMed: 37684529 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-023-05292-1
- Primary Citation Related Structures:
8FWB, 8FWC, 8FWE, 8FWG, 8FWM, 8FXP, 8FXR - PubMed Abstract:
Large gaps exist in our understanding of how bacteriophages, the most abundant biological entities on Earth, assemble and function. The structure of the "neck" region, where the DNA-filled capsid is connected to the host-recognizing tail remains poorly understood. We describe cryo-EM structures of the neck, the neck-capsid and neck-tail junctions, and capsid of the Agrobacterium phage Milano. The Milano neck 1 protein connects the 12-fold symmetrical neck to a 5-fold vertex of the icosahedral capsid. Comparison of Milano neck 1 homologs leads to four proposed classes, likely evolved from the simplest one in siphophages to more complex ones in myo- and podophages. Milano neck is surrounded by the atypical collar, which covalently crosslinks the tail sheath to neck 1. The Milano capsid is decorated with three types of proteins, a minor capsid protein (mCP) and two linking proteins crosslinking the mCP to the major capsid protein. The extensive network of disulfide bonds within and between neck, collar, capsid and tail provides an exceptional structural stability to Milano.
- Department of Biochemistry and Molecular Genetics, University of Virginia School of Medicine, Charlottesville, VA, 22903, USA.
Organizational Affiliation: