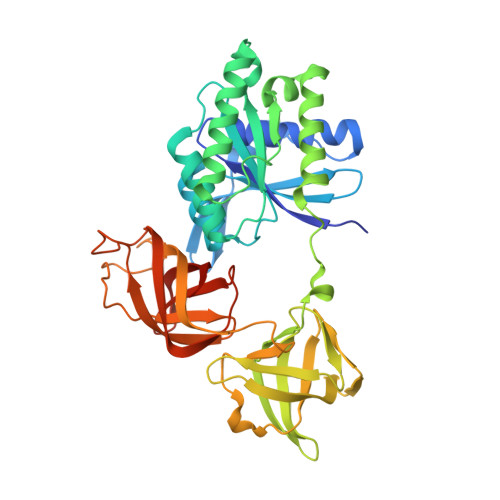
Antibiotic that inhibits trans -translation blocks binding of EF-Tu to tmRNA but not to tRNA.
Marathe, N., Nguyen, H.A., Alumasa, J.N., Kuzmishin Nagy, A.B., Vazquez, M., Dunham, C.M., Keiler, K.C.(2023) mBio 14: e0146123-e0146123
- PubMed: 37681945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.01461-23
- Primary Citation Related Structures:
8FR3 - PubMed Abstract:
Elongation factor thermo-unstable (EF-Tu) is a universally conserved translation factor that mediates productive interactions between tRNAs and the ribosome. In bacteria, EF-Tu also delivers transfer-messenger RNA (tmRNA)-SmpB to the ribosome during trans -translation. We report the first small molecule, KKL-55, that specifically inhibits EF-Tu activity in trans -translation without affecting its activity in normal translation. KKL-55 has broad-spectrum antibiotic activity, suggesting that compounds targeted to the tmRNA-binding interface of EF-Tu could be developed into new antibiotics to treat drug-resistant infections.
- Department of Biochemistry and Molecular Biology, The Pennsylvania State University , University Park, Pennsylvania, USA.
Organizational Affiliation: