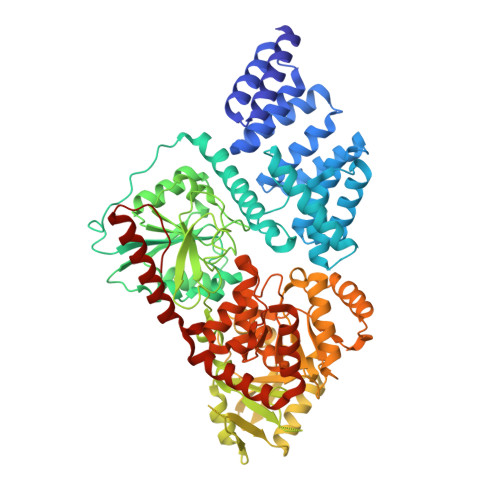
Motif-dependent binding on the intervening domain regulates O-GlcNAc transferase.
Blankenship, C.M., Xie, J., Benz, C., Wang, A., Ivarsson, Y., Jiang, J.(2023) Nat Chem Biol 19: 1423-1431
- PubMed: 37653170 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01422-2
- Primary Citation Related Structures:
8FE6, 8FE7, 8FUF - PubMed Abstract:
The modification of intracellular proteins with O-linked β-N-acetylglucosamine (O-GlcNAc) moieties is a highly dynamic process that spatiotemporally regulates nearly every important cellular program. Despite its significance, little is known about the substrate recognition and regulation modes of O-GlcNAc transferase (OGT), the primary enzyme responsible for O-GlcNAc addition. In this study, we identified the intervening domain (Int-D), a poorly understood protein fold found only in metazoan OGTs, as a specific regulator of OGT protein-protein interactions and substrate modification. Using proteomic peptide phage display (ProP-PD) coupled with structural, biochemical and cellular characterizations, we discovered a strongly enriched peptide motif, employed by the Int-D to facilitate specific O-GlcNAcylation. We further show that disruption of Int-D binding dysregulates important cellular programs, including response to nutrient deprivation and glucose metabolism. These findings illustrate a mode of OGT substrate recognition and offer key insights into the biological roles of this unique domain.
- Pharmaceutical Sciences Division, School of Pharmacy, University of Wisconsin-Madison, Madison, WI, USA.
Organizational Affiliation: