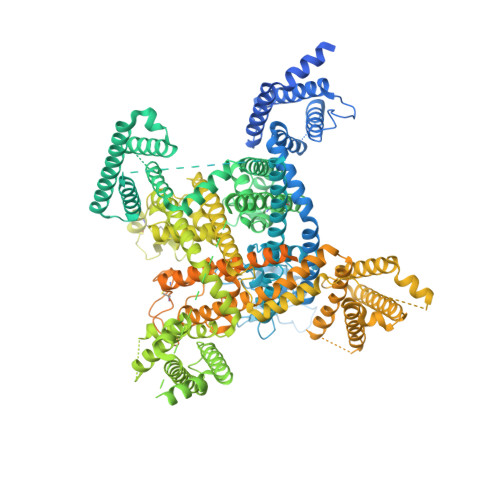
Structural basis for inhibition of the cardiac sodium channel by the atypical antiarrhythmic drug ranolazine.
Lenaeus, M., Gamal El-Din, T.M., Tonggu, L., Zheng, N., Catterall, W.A.(2023) Nat Cardiovasc Res 2: 587-594
- PubMed: 39185478 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44161-023-00271-5
- Primary Citation Related Structures:
8F6P - Division of General Internal Medicine, Department of Medicine, University of Washington, Seattle, WA, USA.
Organizational Affiliation: