Major-groove sequence-specific RNA recognition by LoaP, a paralog of transcription elongation factor NusG.
Elghondakly, A., Jermain, M.D., Winkler, W.C., Ferre-D'Amare, A.R.(2024) Structure 32: 1488
- PubMed: 38959899 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.06.001
- Primary Citation Related Structures:
8F5G - PubMed Abstract:
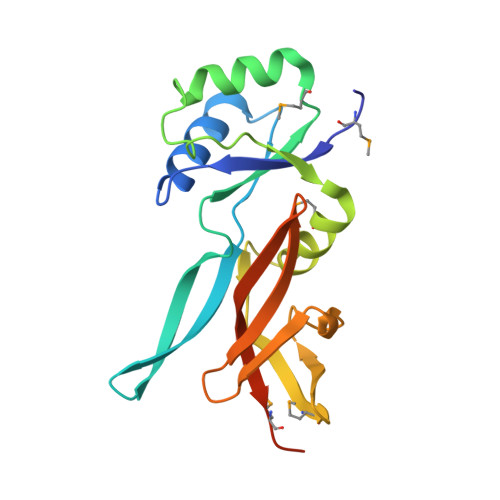
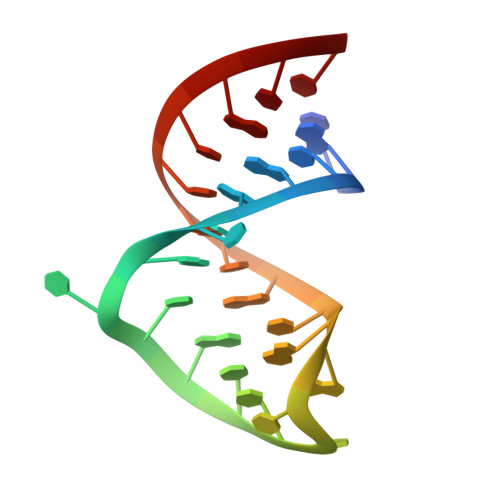
LoaP is a member of the universal NusG protein family. Previously, we reported that unlike other characterized homologs, LoaP binds RNA sequence-specifically, recognizing a stem-loop in the 5'-untranslated region of operons it regulates. To elucidate how this NusG homolog acquired this ability, we now determined the co-crystal structure of Thermoanaerobacter pseudethanolicus LoaP bound to its cognate 26-nucleotide dfn RNA element. Our structure reveals that the LoaP C-terminal KOW domain recognizes the helical portion of the RNA by docking into a broadened major groove, while a protruding β-hairpin of the N-terminal NusG-like domain binds the UNCG tetraloop capping the stem-loop. Major-groove RNA recognition is unusual and is made possible by conserved features of the dfn hairpin. Superposition with structures of other NusG proteins implies that LoaP can bind concurrently to the dfn RNA and the transcription elongation complex, suggesting a new level of co-transcriptional regulation by proteins of this conserved family.
- Laboratory of Nucleic Acids, National Heart, Lung, and Blood Institute, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: