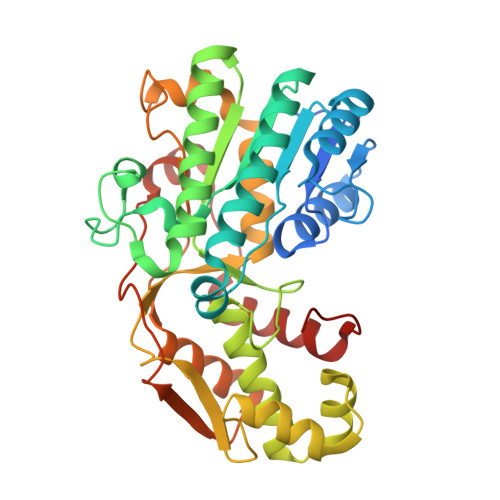
Bifunctional Epimerase/Reductase Enzymes Facilitate the Modulation of 6-Deoxy-Heptoses Found in the Capsular Polysaccharides of Campylobacter jejuni.
Xiang, D.F., Ghosh, M.K., Riegert, A.S., Thoden, J.B., Holden, H.M., Raushel, F.M.(2023) Biochemistry 62: 134-144
- PubMed: 36534477 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.2c00633
- Primary Citation Related Structures:
8EWU - PubMed Abstract:
Campylobacter jejuni is a human pathogen and the leading cause of food poisoning in the United States and Europe. Surrounding the exterior surface of this bacterium is a capsular polysaccharide (CPS) that consists of a repeating sequence of common and unusual carbohydrate segments. At least 10 different heptose sugars have thus far been identified in the various strains of C. jejuni . The accepted biosynthetic pathway for the construction of the 6-deoxy-heptoses begins with the 4,6-dehydration of GDP-d- glycero -d- manno -heptose by a dehydratase, followed by an epimerase that racemizes C3 and/or C5 of the product GDP-6-deoxy-4-keto-d- lyxo -heptose. In the final step, a C4-reductase catalyzes the NADPH reduction of the resulting 4-keto product. However, in some strains and serotypes of C. jejuni , there are two separate C4-reductases with different product specificities in the gene cluster for CPS formation. Five pairs of these tandem C4-reductases were isolated, and the catalytic properties were ascertained. In four out of five cases, one of the two C4-reductases is able to catalyze the isomerization of C3 and C5 of GDP-6-deoxy-4-keto-d- lyxo -heptose, in addition to the catalysis of the reduction of C4, thus bypassing the requirement for a separate C3/C5-isomerase. In each case, the 3'-end of the gene for the first C4-reductase contains a poly-G tract of 8-10 guanine residues that may be used to control the expression and/or catalytic activity of either C4-reductase. The three-dimensional structure of the C4-reductase from serotype HS:15, which only does a reduction of C4, was determined to 1.45 Å resolution in the presence of NADPH and GDP.
- Department of Chemistry, Texas A&M University, College Station, Texas 77843, United States.
Organizational Affiliation: