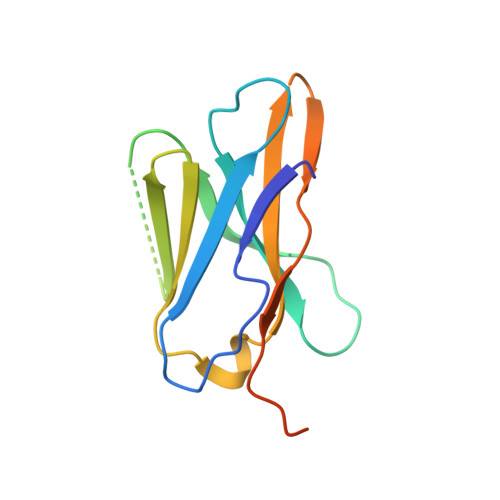
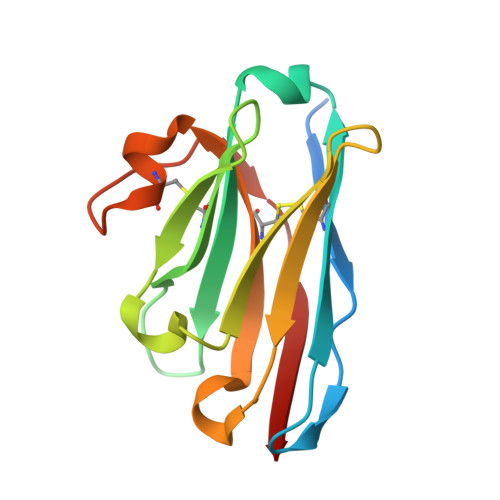
Development of an 18 F-labeled anti-human CD8 VHH for same-day immunoPET imaging.
Sriraman, S.K., Davies, C.W., Gill, H., Kiefer, J.R., Yin, J., Ogasawara, A., Urrutia, A., Javinal, V., Lin, Z., Seshasayee, D., Abraham, R., Haas, P., Koth, C., Marik, J., Koerber, J.T., Williams, S.P.(2023) Eur J Nucl Med Mol Imaging 50: 679-691
- PubMed: 36346438 Search on PubMed
- DOI: https://doi.org/10.1007/s00259-022-05998-0
- Primary Citation Related Structures:
8EW6 - PubMed Abstract:
Cancer immunotherapies (CITs) have revolutionized the treatment of certain cancers, but many patients fail to respond or relapse from current therapies, prompting the need for new CIT agents. CD8 + T cells play a central role in the activity of many CITs, and thus, the rapid imaging of CD8 + cells could provide a critical biomarker for new CIT agents. However, existing 89 Zr-labeled CD8 PET imaging reagents exhibit a long circulatory half-life and high radiation burden that limit potential applications such as same-day and longitudinal imaging. To this end, we discovered and developed a 13-kDa single-domain antibody (VHH5v2) against human CD8 to enable high-quality, same-day imaging with a reduced radiation burden. To enable sensitive and rapid imaging, we employed a site-specific conjugation strategy to introduce an 18 F radiolabel to the VHH. The anti-CD8 VHH, VHH5v2, demonstrated binding to a membrane distal epitope of human CD8 with a binding affinity (K D ) of 500 pM. Subsequent imaging experiments in several xenografts that express varying levels of CD8 demonstrated rapid tumor uptake and fast clearance from the blood. High-quality images were obtained within 1 h post-injection and could quantitatively differentiate the tumor models based on CD8 expression level. Our work reveals the potential of this anti-human CD8 VHH [ 18 F]F-VHH5v2 to enable rapid and specific imaging of CD8 + cells in the clinic.
- Department of Biomedical Imaging, Genentech, Inc, 1 DNA Way, South San Francisco, Genetech, CA, 94080, USA.
Organizational Affiliation: