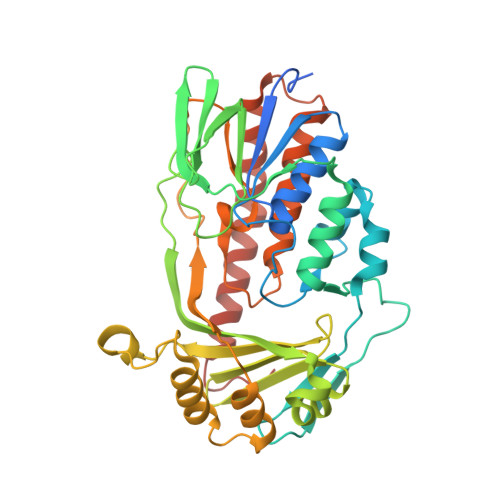
Structure of anhydrotetracycline-bound Tet(X6) reveals the mechanism for inhibition of type 1 tetracycline destructases.
Kumar, H., Williford, E.E., Blake, K.S., Virgin-Downey, B., Dantas, G., Wencewicz, T.A., Tolia, N.H.(2023) Commun Biol 6: 423-423
- PubMed: 37062778 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-023-04792-4
- Primary Citation Related Structures:
8ER0, 8ER1 - PubMed Abstract:
Inactivation of tetracycline antibiotics by tetracycline destructases (TDases) remains a clinical and agricultural threat. TDases can be classified as type 1 Tet(X)-like TDases and type 2 soil-derived TDases. Type 1 TDases are widely identified in clinical pathogens. A combination therapy of tetracycline and a TDase inhibitor is much needed to rescue the clinical efficacy of tetracyclines. Anhydrotetracycline is a pan-TDase inhibitor that inhibits both type 1 and type 2 TDases. Here, we present structural, biochemical, and phenotypic evidence that anhydrotetracycline binds in a substrate-like orientation and competitively inhibits the type 1 TDase Tet(X6) to rescue tetracycline antibiotic activity as a sacrificial substrate. Anhydrotetracycline interacting residues of Tet(X6) are conserved within type 1 TDases, indicating a conserved binding mode and mechanism of inhibition. This mode of binding and inhibition is distinct from anhydrotetracycline's inhibition of type 2 TDases. This study forms the framework for development of next-generation therapies to counteract enzymatic tetracycline resistance.
- Host-pathogen interaction and structural vaccinology section (HPISV), National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (NIH), Bethesda, MD, USA.
Organizational Affiliation: