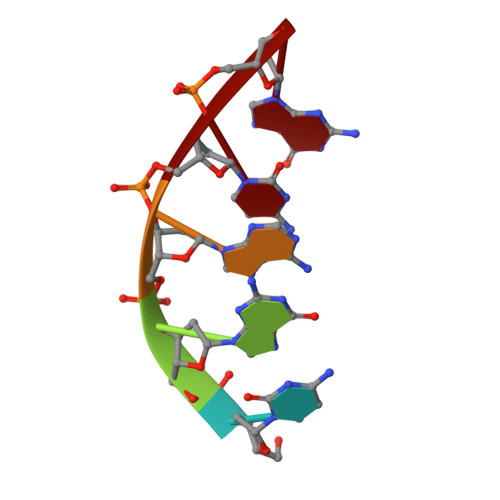
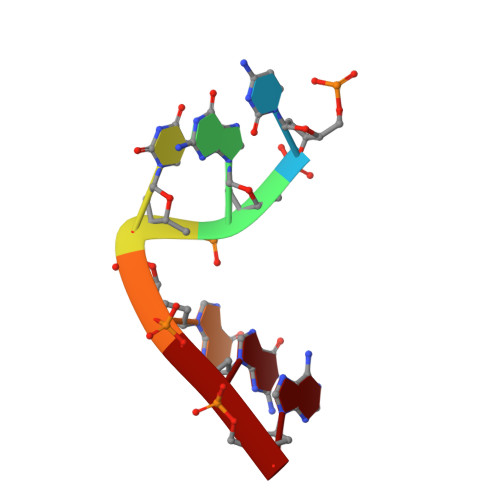
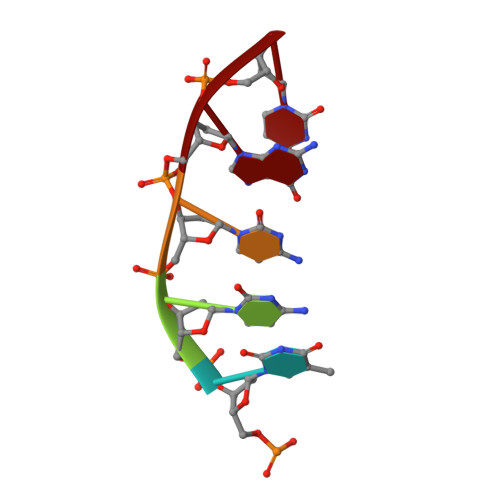
Engineering DNA Crystals toward Studying DNA-Guest Molecule Interactions.
Zhang, C., Zhao, J., Lu, B., Seeman, N.C., Sha, R., Noinaj, N., Mao, C.(2023) J Am Chem Soc 145: 4853-4859
- PubMed: 36791277 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.3c00081
- Primary Citation Related Structures:
8EP8, 8EPB, 8EPD, 8EPE, 8EPF, 8EPG, 8EPI, 8F40, 8F42 - PubMed Abstract:
Sequence-selective recognition of DNA duplexes is important for a wide range of applications including regulating gene expression, drug development, and genome editing. Many small molecules can bind DNA duplexes with sequence selectivity. It remains as a challenge how to reliably and conveniently obtain the detailed structural information on DNA-molecule interactions because such information is critically needed for understanding the underlying rules of DNA-molecule interactions. If those rules were understood, we could design molecules to recognize DNA duplexes with a sequence preference and intervene in related biological processes, such as disease treatment. Here, we have demonstrated that DNA crystal engineering is a potential solution. A molecule-binding DNA sequence is engineered to self-assemble into highly ordered DNA crystals. An X-ray crystallographic study of molecule-DNA cocrystals reveals the structural details on how the molecule interacts with the DNA duplex. In this approach, the DNA will serve two functions: (1) being part of the molecule to be studied and (2) forming the crystal lattice. It is conceivable that this method will be a general method for studying drug/peptide-DNA interactions. The resulting DNA crystals may also find use as separation matrices, as hosts for catalysts, and as media for material storage.
- Department of Chemistry, Purdue University, West Lafayette, Indiana 47907, United States.
Organizational Affiliation: