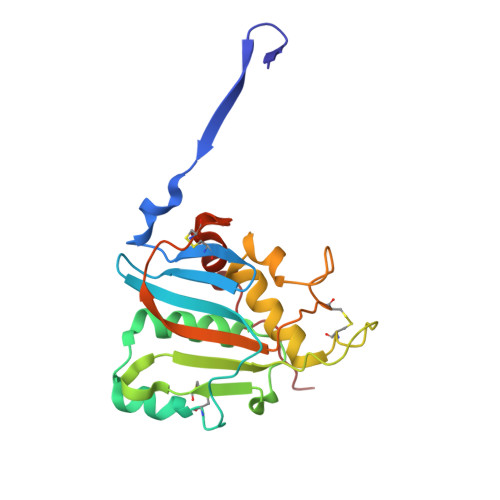
Structural conservation of Lassa virus glycoproteins and recognition by neutralizing antibodies.
Perrett, H.R., Brouwer, P.J.M., Hurtado, J., Newby, M.L., Liu, L., Muller-Krauter, H., Muller Aguirre, S., Burger, J.A., Bouhuijs, J.H., Gibson, G., Messmer, T., Schieffelin, J.S., Antanasijevic, A., Boons, G.J., Strecker, T., Crispin, M., Sanders, R.W., Briney, B., Ward, A.B.(2023) Cell Rep 42: 112524-112524
- PubMed: 37209096 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2023.112524
- Primary Citation Related Structures:
8EJD, 8EJE, 8EJF, 8EJG, 8EJH, 8EJI, 8EJJ - PubMed Abstract:
Lassa fever is an acute hemorrhagic fever caused by the zoonotic Lassa virus (LASV). The LASV glycoprotein complex (GPC) mediates viral entry and is the sole target for neutralizing antibodies. Immunogen design is complicated by the metastable nature of recombinant GPCs and the antigenic differences among phylogenetically distinct LASV lineages. Despite the sequence diversity of the GPC, structures of most lineages are lacking. We present the development and characterization of prefusion-stabilized, trimeric GPCs of LASV lineages II, V, and VII, revealing structural conservation despite sequence diversity. High-resolution structures and biophysical characterization of the GPC in complex with GP1-A-specific antibodies suggest their neutralization mechanisms. Finally, we present the isolation and characterization of a trimer-preferring neutralizing antibody belonging to the GPC-B competition group with an epitope that spans adjacent protomers and includes the fusion peptide. Our work provides molecular detail information on LASV antigenic diversity and will guide efforts to design pan-LASV vaccines.
- Department of Integrative, Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: