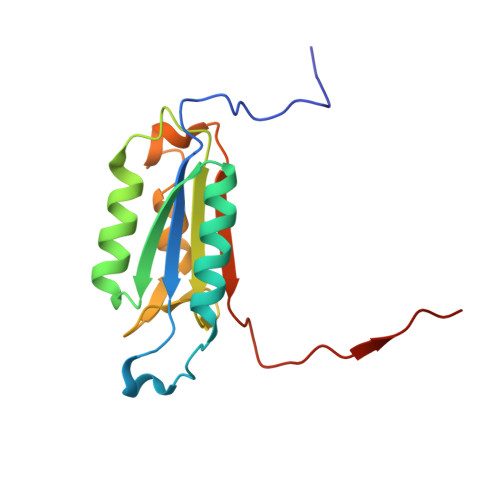
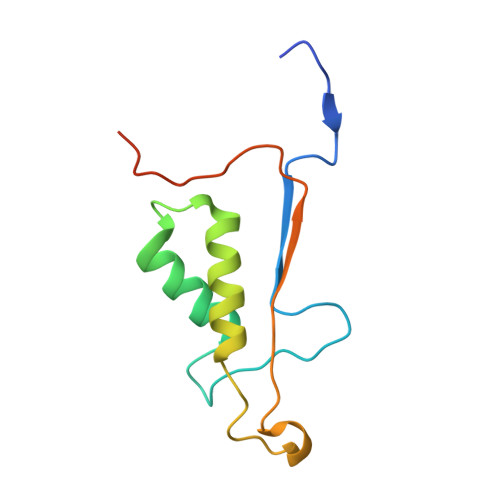
Engaging a Non-catalytic Cysteine Residue Drives Potent and Selective Inhibition of Caspase-6.
Van Horn, K.S., Wang, D., Medina-Cleghorn, D., Lee, P.S., Bryant, C., Altobelli, C., Jaishankar, P., Leung, K.K., Ng, R.A., Ambrose, A.J., Tang, Y., Arkin, M.R., Renslo, A.R.(2023) J Am Chem Soc 145: 10015-10021
- PubMed: 37104712 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.2c12240
- Primary Citation Related Structures:
8EG5, 8EG6 - PubMed Abstract:
Caspases are a family of cysteine-dependent proteases with important cellular functions in inflammation and apoptosis, while also implicated in human diseases. Classical chemical tools to study caspase functions lack selectivity for specific caspase family members due to highly conserved active sites and catalytic machinery. To overcome this limitation, we targeted a non-catalytic cysteine residue (C264) unique to caspase-6 (C6), an enigmatic and understudied caspase isoform. Starting from disulfide ligands identified in a cysteine trapping screen, we used a structure-informed covalent ligand design to produce potent, irreversible inhibitors ( 3a ) and chemoproteomic probes ( 13- t ) of C6 that exhibit unprecedented selectivity over other caspase family members and high proteome selectivity. This approach and the new tools described will enable rigorous interrogation of the role of caspase-6 in developmental biology and in inflammatory and neurodegenerative diseases.
- Department of Pharmaceutical Chemistry, University of California, San Francisco, 600 16th Street, San Francisco, California 94143, United States.
Organizational Affiliation: