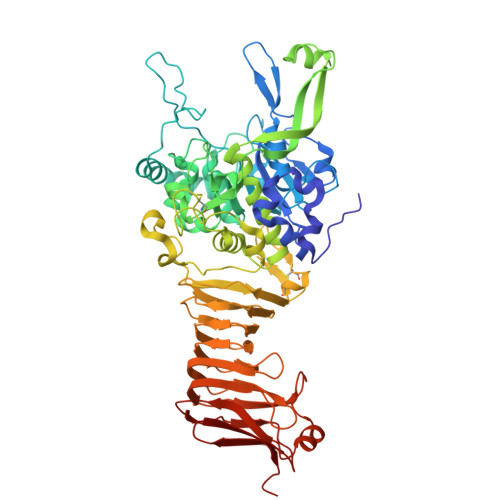
Crystal structure of a subtilisin-like autotransporter passenger domain reveals insights into its cytotoxic function.
Hor, L., Pilapitiya, A., McKenna, J.A., Panjikar, S., Anderson, M.A., Desvaux, M., Paxman, J.J., Heras, B.(2023) Nat Commun 14: 1163-1163
- PubMed: 36859523 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-36719-2
- Primary Citation Related Structures:
8E7F - PubMed Abstract:
Autotransporters (ATs) are a large family of bacterial secreted and outer membrane proteins that encompass a wide range of enzymatic activities frequently associated with pathogenic phenotypes. We present the structural and functional characterisation of a subtilase autotransporter, Ssp, from the opportunistic pathogen Serratia marcescens. Although the structures of subtilases have been well documented, this subtilisin-like protein is associated with a 248 residue β-helix and itself includes three finger-like protrusions around its active site involved in substrate interactions. We further reveal that the activity of the subtilase AT is required for entry into epithelial cells as well as causing cellular toxicity. The Ssp structure not only provides details about the subtilase ATs, but also reveals a common framework and function to more distantly related ATs. As such these findings also represent a significant step forward toward understanding the molecular mechanisms underlying the functional divergence in the large AT superfamily.
- Department of Biochemistry and Chemistry, La Trobe Institute for Molecular Science, La Trobe University, Kingsbury Drive, Bundoora, VIC, 3086, Australia.
Organizational Affiliation: