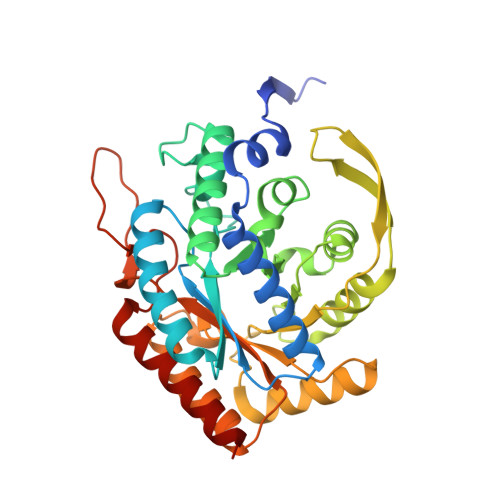
Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Balachandran, N., Grainger, R.A., Rob, T., Liuni, P., Wilson, D.J., Junop, M.S., Berti, P.J.(2022) Biochemistry 61: 2229-2240
- PubMed: 36197914 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.2c00465
- Primary Citation Related Structures:
8E0S, 8E0T, 8E0U, 8E0V, 8E0X, 8E0Y, 8E0Z - PubMed Abstract:
α-Carboxyketose synthases, including 3-deoxy-d- arabino heptulosonate 7-phosphate synthase (DAHPS), are long-standing targets for inhibition. They are challenging targets to create tight-binding inhibitors against, and inhibitors often display half-of-sites binding and partial inhibition. Half-of-sites inhibition demonstrates the existence of inter-subunit communication in DAHPS. We used X-ray crystallography and spatially resolved hydrogen-deuterium exchange (HDX) to reveal the structural and dynamic bases for inter-subunit communication in Escherichia coli DAHPS(Phe), the isozyme that is feedback-inhibited by phenylalanine. Crystal structures of this homotetrameric (dimer-of-dimers) enzyme are invariant over 91% of its sequence. Three variable loops make up 8% of the sequence and are all involved in inter-subunit contacts across the tight-dimer interface. The structures have pseudo-twofold symmetry indicative of inter-subunit communication across the loose-dimer interface, with the diagonal subunits B and C always having the same conformation as each other, while subunits A and D are variable. Spatially resolved HDX reveals contrasting responses to ligand binding, which, in turn, affect binding of the second substrate, erythrose-4-phosphate (E4P). The N -terminal peptide, M1-E12, and the active site loop that binds E4P, F95-K105, are key parts of the communication network. Inter-subunit communication appears to have a catalytic role in all α-carboxyketose synthase families and a regulatory role in some members.
- Department of Biochemistry, Molecular Biology Lab, Western University, London, Ontario N6A 5C1, Canada.
Organizational Affiliation: