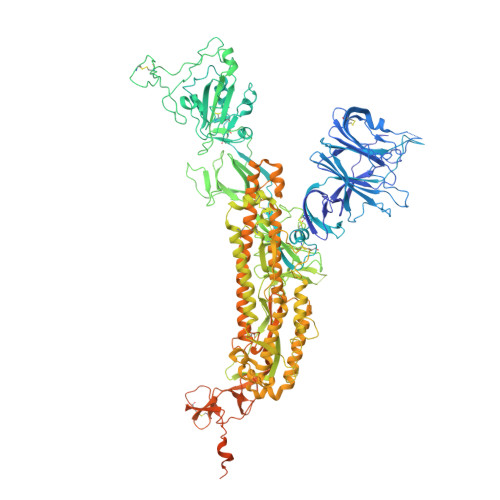
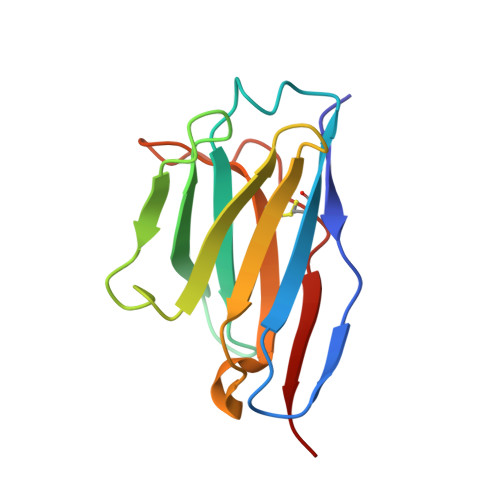
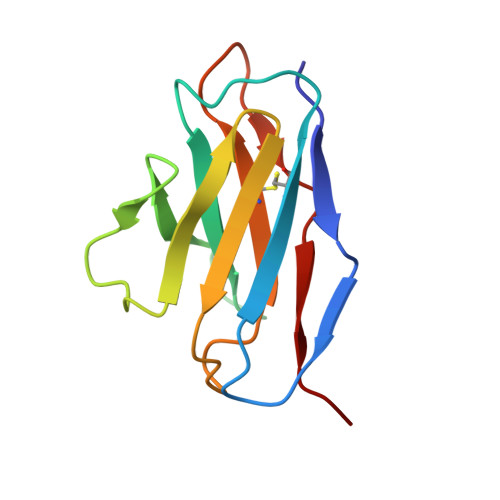
Function and Cryo-EM structures of broadly potent bispecific antibodies against multiple SARS-CoV-2 Omicron sublineages.
Ren, P., Hu, Y., Peng, L., Yang, L., Suzuki, K., Fang, Z., Bai, M., Zhou, L., Feng, Y., Zou, Y., Xiong, Y., Chen, S.(2023) Signal Transduct Target Ther 8: 281-281
- PubMed: 37518189 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41392-023-01509-1
- Primary Citation Related Structures:
8DZH, 8DZI - Department of Genetics, Yale University School of Medicine, New Haven, CT, USA.
Organizational Affiliation: