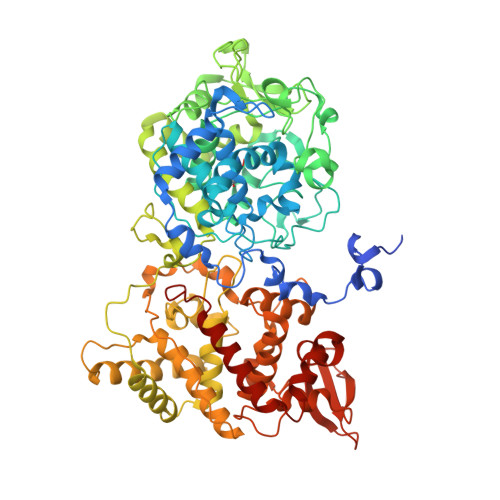
Characterization of a catalase-peroxidase variant (L333V-KatG) identified in an INH-resistant Mycobacterium tuberculosis clinical isolate.
Uribe-Vazquez, B., Diaz-Vilchis, A., Avila-Linares, A., Saab-Rincon, G., Marin-Tovar, Y., Flores, H., Pastor, N., Huerta-Miranda, G., Rudino-Pinera, E., Soberon, X.(2024) Biochem Biophys Rep 37: 101649-101649
- PubMed: 38318524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrep.2024.101649
- Primary Citation Related Structures:
8DWR - PubMed Abstract:
Mycobacterium tuberculosis catalase-peroxidase ( Mt -KatG) is a bifunctional heme-dependent enzyme that has been shown to activate isoniazid (INH), the widely used antibiotic against tuberculosis (TB). The L333V-KatG variant has been associated with INH resistance in clinical M. tuberculosis isolates from Mexico. To understand better the mechanisms of INH activation, its catalytic properties (catalase, peroxidase, and IN-NAD formation) and crystal structure were compared with those of the wild-type enzyme (WT-KatG). The rate of IN-NAD formation mediated by WT-KatG was 23% greater than L333V-KatG when INH concentration is varied. In contrast to WT-KatG, the crystal structure of the L333V-KatG variant has a perhydroxy modification of the indole nitrogen of W107 from MYW adduct. L333V-KatG shows most of the active site residues in a similar position to WT-KatG; only R418 is in the R-conformation instead of the double R and Y conformation present in WT-KatG. L333V-KatG shows a small displacement respect to WT-KatG in the helix from R385 to L404 towards the mutation site, an increase in length of the coordination bond between H270 and heme Fe, and a longer H-bond between proximal D381 and W321, compared to WT-KatG; these small displacements could explain the altered redox potential of the heme, and result in a less active and stable enzyme.
- Departamento de Ingeniería Celular y Biocatálisis, Instituto de Biotecnología, UNAM, Avenida Universidad 2001, Colonia Chamilpa, 62210, Cuernavaca, México.
Organizational Affiliation: