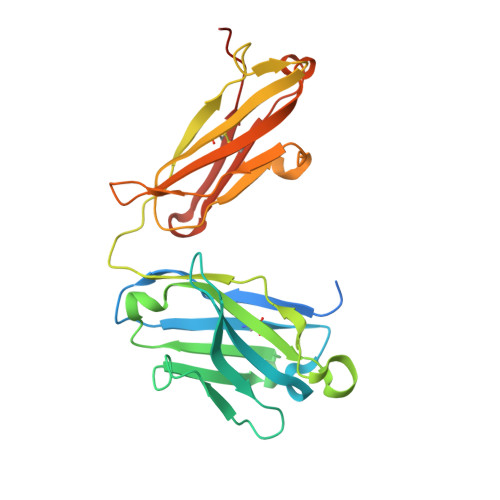
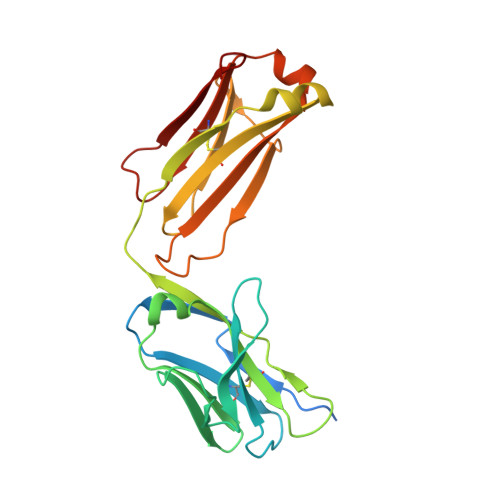
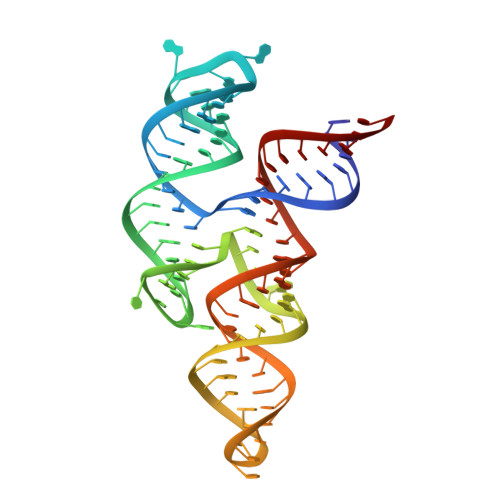
Crystal structure of a highly conserved enteroviral 5' cloverleaf RNA replication element.
Das, N.K., Hollmann, N.M., Vogt, J., Sevdalis, S.E., Banna, H.A., Ojha, M., Koirala, D.(2023) Nat Commun 14: 1955-1955
- PubMed: 37029118 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-37658-8
- Primary Citation Related Structures:
8DP3 - PubMed Abstract:
The extreme 5'-end of the enterovirus RNA genome contains a conserved cloverleaf-like domain that recruits 3CD and PCBP proteins required for initiating genome replication. Here, we report the crystal structure at 1.9 Å resolution of this domain from the CVB3 genome in complex with an antibody chaperone. The RNA folds into an antiparallel H-type four-way junction comprising four subdomains with co-axially stacked sA-sD and sB-sC helices. Long-range interactions between a conserved A40 in the sC-loop and Py-Py helix within the sD subdomain organize near-parallel orientations of the sA-sB and sC-sD helices. Our NMR studies confirm that these long-range interactions occur in solution and without the chaperone. The phylogenetic analyses indicate that our crystal structure represents a conserved architecture of enteroviral cloverleaf-like domains, including the A40 and Py-Py interactions. The protein binding studies further suggest that the H-shape architecture provides a ready-made platform to recruit 3CD and PCBP2 for viral replication.
- Department of Chemistry and Biochemistry, University of Maryland Baltimore County, Baltimore, MD, 21250, USA.
Organizational Affiliation: